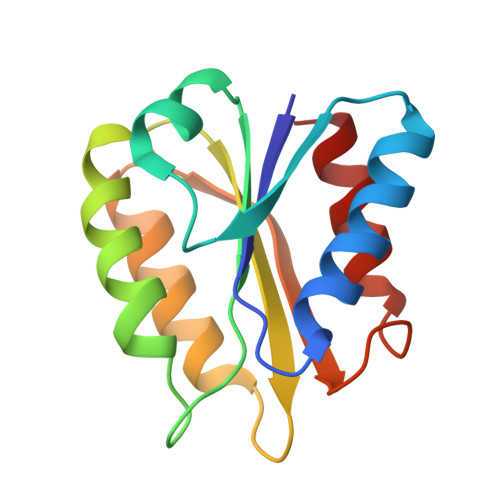
Tertiary structure of two-electron reduced Megasphaera elsdenii flavodoxin and some implications, as determined by two-dimensional 1H NMR and restrained molecular dynamics
van Mierlo, C.P.M., Lijnzaad, P., Vervoort, J., Mueller, F., Berendsen, H.J., de Vlieg, J.(1990) Eur J Biochem 194: 185-198
- PubMed: 2253614 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1990.tb19444.x
- Primary Citation Related Structures:
2FZ5 - PubMed Abstract:
The tertiary structure of the non-crystallizable two-electron-reduced Megasphaera elsdenii flavodoxin (15 kDa, 137 amino acid residues) has been determined using nuclear Overhauser enhancement restraints extracted from two-dimensional 1H-NMR spectra. A tertiary structure satisfying the experimental restraints very well (maximum NOE violation of 66 pm) was obtained with use of restrained molecular dynamics, using 509 distance restraints (including one non-NOE) on a starting structure modeled from the crystal structure of one-electron-reduced Clostridium MP flavodoxin. The protein consists of a central parallel beta-sheet surrounded on both sides by two alpha-helices. The flavin is positioned at the periphery of the molecule. The tertiary structure of the protein is highly defined with the exception of the flavin. The latter is expected to result from performing the restrained molecular dynamics simulation without water molecules and without proper charges on the flavin. The flavin, including the phosphate, the ribityl side chain and the isoalloxazine ring, is solvent accessible under the experimental conditions used and evidenced by a two-dimensional amide exchange experiment. This accessibility is expected to be important in the redox potential regulation of the semiquinone/hydroquinone couple of the protein. The amide exchange against deuterons and several typical line shapes in the two-dimensional NMR spectra are consistent with the structure generated. The structure is discussed in detail.
- Department of Biochemistry, Agricultural University, Wageningen, The Netherlands.
Organizational Affiliation: