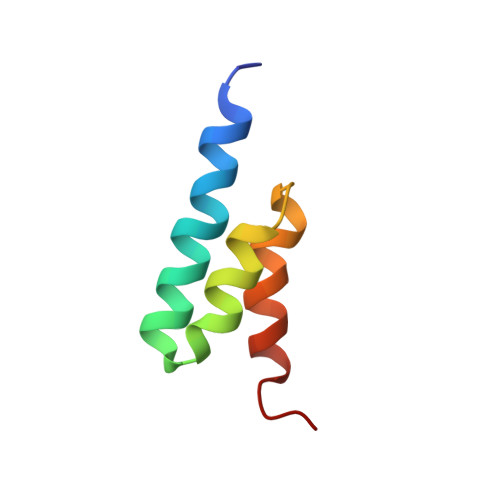
Structure, dynamics, and stability variation in bacterial albumin binding modules: implications for species specificity.
He, Y., Rozak, D.A., Sari, N., Chen, Y., Bryan, P., Orban, J.(2006) Biochemistry 45: 10102-10109
- PubMed: 16906768 Search on PubMed
- DOI: https://doi.org/10.1021/bi060409m
- Primary Citation Related Structures:
2FS1 - PubMed Abstract:
Protein G-related albumin-binding (GA) modules are frequently expressed on the surfaces of bacterial cells. The limited amino acid sequence variation among GA modules results in structural and functional differences with possible implications for bacterial pathogenesis and host specificity. In particular, the streptococcal G148-GA3 and F. magna ALB8-GA albumin-binding domains exhibit a degree of structural and dynamic diversity that may account for their varied affinities for different species of albumin. To explore the impact of GA module polymorphisms on albumin binding and specificity, we recently used offset recombinant PCR to shuffle seven artificially constructed representatives of the GA sequence space and scan the phage-displayed recombinant domains for mutations that supported binding to the phylogenetically distinct human and guinea pig serum albumins (HSA and GPSA) (Rozak et al. (2006) Biochemistry 45, 3263-3271). Surprisingly, phage selection revealed an overwhelming preference for a single recombinant domain (PSD-1, phage-selected domain-1) regardless of whether the phages were enriched for their abilities to bind one or both of these albumins. We describe here the NMR-derived structure, dynamics, and stability of unbound PSD-1. Our results demonstrate that increased flexibility is not a requirement for broadened specificity, as had been suggested earlier (Johansson et al. (2002) J. Mol. Biol. 316, 1083-1099), because PSD-1 binds the phylogenetically diverse HSA and GPSA even more tightly than G148-GA3 but is less flexible. The structural basis for albumin-binding specificity is analyzed in light of these new results.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, Maryland 20850, USA.
Organizational Affiliation: