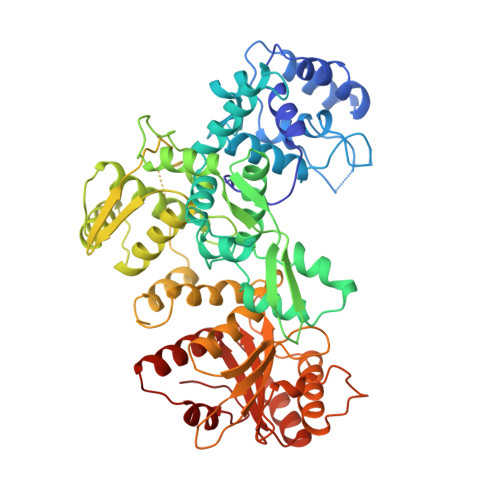
Structure of FokI has implications for DNA cleavage.
Wah, D.A., Bitinaite, J., Schildkraut, I., Aggarwal, A.K.(1998) Proc Natl Acad Sci U S A 95: 10564-10569
- PubMed: 9724743 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.95.18.10564
- Primary Citation Related Structures:
2FOK - PubMed Abstract:
FokI is a member an unusual class of restriction enzymes that recognize a specific DNA sequence and cleave nonspecifically a short distance away from that sequence. FokI consists of an N-terminal DNA recognition domain and a C-terminal cleavage domain. The bipartite nature of FokI has led to the development of artificial enzymes with novel specificities. We have solved the structure of FokI to 2.3 A resolution. The structure reveals a dimer, in which the dimerization interface is mediated by the cleavage domain. Each monomer has an overall conformation similar to that found in the FokI-DNA complex, with the cleavage domain packing alongside the DNA recognition domain. In corroboration with the cleavage data presented in the accompanying paper in this issue of Proceedings, we propose a model for FokI DNA cleavage that requires the dimerization of FokI on DNA to cleave both DNA strands.
- Structural Biology Program, Department of Physiology and Biophysics, Box 1677, 1425 Madison Avenue, Mount Sinai School of Medicine, New York, NY 10029, USA.
Organizational Affiliation: