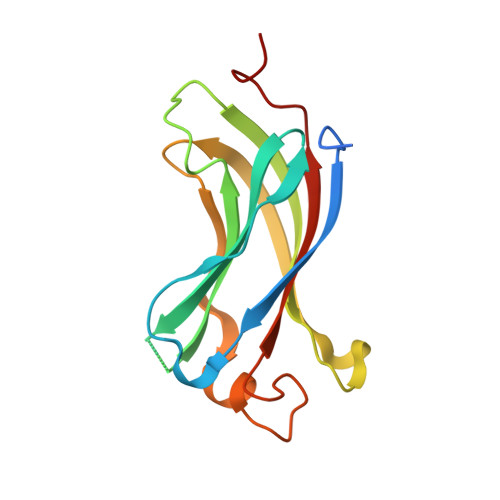
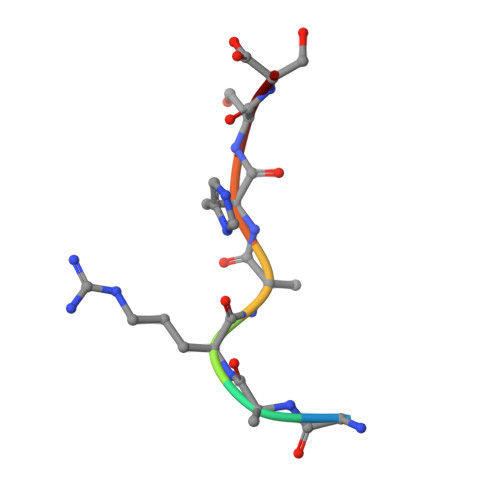
Molecular recognition of p53 and MDM2 by USP7/HAUSP
Sheng, Y., Saridakis, V., Sarkari, F., Duan, S., Wu, T., Arrowsmith, C.H., Frappier, L.(2006) Nat Struct Mol Biol 13: 285-291
- PubMed: 16474402 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1067
- Primary Citation Related Structures:
2FOJ, 2FOO, 2FOP - PubMed Abstract:
The ubiquitin-specific protease, USP7, has key roles in the p53 pathway whereby it stabilizes both p53 and MDM2. We show that the N-terminal domain of USP7 binds two closely spaced 4-residue sites in both p53 and MDM2, falling between p53 residues 359-367 and MDM2 residues 147-159. Cocrystal structures with USP7 were determined for both p53 peptides and for one MDM2 peptide. These peptides bind the same surface of USP7 as Epstein-Barr nuclear antigen-1, explaining the competitive nature of the interactions. The structures and mutagenesis data indicate a preference for a P/AXXS motif in peptides that bind USP7. Contacts made by serine are identical and crucial for all peptides, and Trp165 in the peptide-binding pocket of USP7 is also crucial. These results help to elucidate the mechanism of substrate recognition by USP7 and the regulation of the p53 pathway.
- Ontario Cancer Institute, Department of Medical Biophysics, 101 College Street, Toronto, Ontario, Canada M5G 1L7.
Organizational Affiliation: