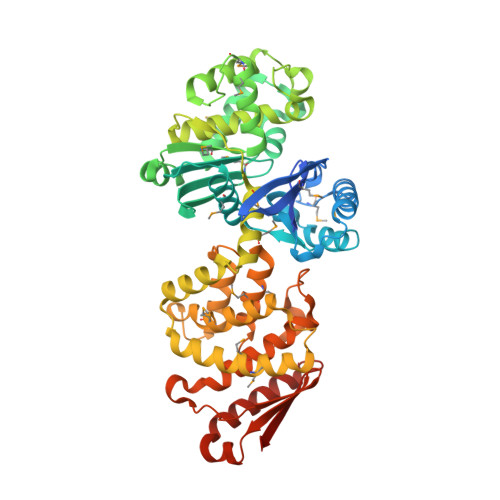
The Structure of the Exopolyphosphatase (PPX) from Escherichia coli O157:H7 Suggests a Binding Mode for Long Polyphosphate Chains.
Rangarajan, E.S., Nadeau, G., Li, Y., Wagner, J., Hung, M.N., Schrag, J.D., Cygler, M., Matte, A.(2006) J Mol Biology 359: 1249-1260
- PubMed: 16678853
- DOI: https://doi.org/10.1016/j.jmb.2006.04.031
- Primary Citation Related Structures:
2FLO - PubMed Abstract:
Polyphosphate (polyP) is a linear polymer consisting of tens to hundreds of phosphate molecules joined together by high-energy anhydride bonds. These polymers are found in virtually all prokaryotic and eukaryotic cells and perform many functions; prominent among them are the responses to many stresses. Polyphosphate is synthesized by polyP kinase (PPK), using the terminal phosphate of ATP as the substrate, and degraded to inorganic phosphate by both endo- and exopolyphosphatases. Here we report the crystal structure and analysis of the polyphosphate phosphatase PPX from Escherichia coli O157:H7 refined at 2.2 Angstroms resolution. PPX is made of four domains. Domains I and II display structural similarity with one another and share the ribonuclease-H-like fold. Domain III bears structural similarity to the N-terminal, HD domain of SpoT. Domain IV, the smallest domain, has structural counterparts in cold-shock associated RNA-binding proteins but is of unknown function in PPX. The putative PPX active site is located at the interface between domains I and II. In the crystal structure of PPX these two domains are close together and represent the "closed" state. Comparison with the crystal structure of PPX/GPPA from Aquifex aeolicus reveals close structural similarity between domains I and II of the two enzymes, with the PPX/GPPA representing an "open" state. A striking feature of the dimer is a deep S-shaped canyon extending along the dimer interface and lined with positively charged residues. The active site region opens to this canyon. We postulate that this is a likely site of polyP binding.
- Department of Biochemistry, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: