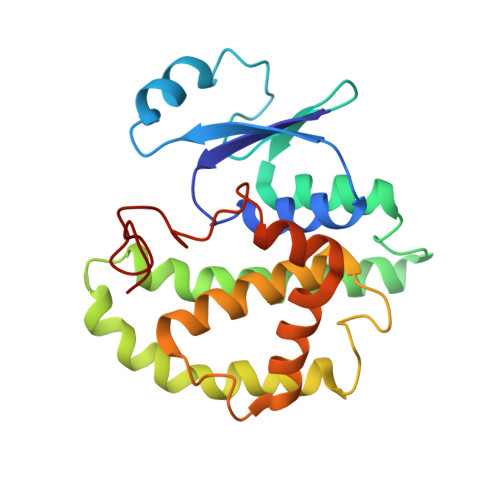
Crystallization, structural determination and analysis of a novel parasite vaccine candidate: Fasciola hepatica glutathione S-transferase.
Rossjohn, J., Feil, S.C., Wilce, M.C., Sexton, J.L., Spithill, T.W., Parker, M.W.(1997) J Mol Biology 273: 857-872
- PubMed: 9367777 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1338
- Primary Citation Related Structures:
1FHE, 2FHE - PubMed Abstract:
Glutathione S-transferases (GSTs) represent the major class of detoxifying enzymes from parasitic helminths. As a result, they are candidates for chemotherapeutic and vaccine design. Indeed, GSTs from Fasciola hepatica have been found to be effective for vaccinating sheep and cattle against fasciolosis. This helminth contains at least seven GST isoforms, of which four have been cloned. The cloned isoforms (Fh51, Fh47, Fh7 and Fh1) all belong to the mu class of GSTs, share greater than 71% sequence identity, yet display distinct substrate specificities. Crystals of Fh47 were obtained using the hanging drop vapour diffusion technique. The crystals belong to space group I4122, with one monomer in the asymmetric unit, which corresponds to a very high solvent content of approximately 75%. The physiological dimer is generated via a crystallographic 2-fold rotation. The three-dimensional structure of Fh47 was solved by molecular replacement using the Schistosoma japonicum glutathione S-transferase (Sj26) crystal structure as a search model. The structure adopts the canonical GST fold comprising two domains: an N-terminal glutathione-binding domain, consisting of a four-stranded beta-sheet and three helices whilst the C-terminal domain is entirely alpha-helical. The presence of Phe19 in Fh47 results in a 6 degrees interdomain rotation in comparison to Sj26, where the equivalent residue is a leucine. Homology models of Fh51, Fh7 and Fh1, based on the Fh47 crystal structure, reveal critical differences in the residues lining the xenobiotic binding site, particularly at residue positions 9, 106 and 204. In addition, differences amongst the isoforms in the non-substrate binding site were noted, which may explain the observed differential binding of large ligands. The major immunogenic epitopes of Fh47 were surprisingly found not to reside on the most solvent-exposed regions of the molecule.
- The Ian Potter Foundation Protein Crystallography Laboratory, St Vincent's Institute of Medical Research, 41 Victoria Parade, Fitzroy, Victoria, 3065, Australia.
Organizational Affiliation: