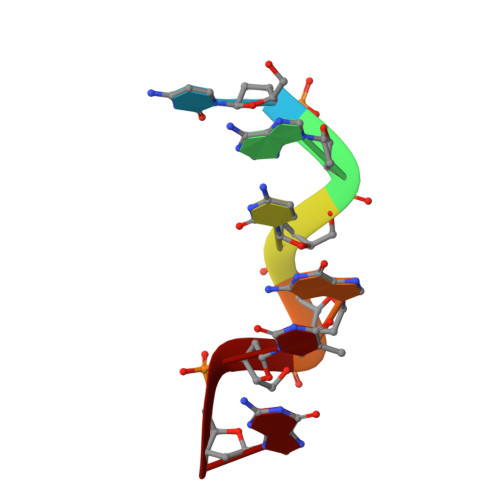
Interaction between the Z-Type DNA Duplex and 1,3-Propanediamine: Crystal Structure of d(CACGTG)2 at 1.2 A Resolution
Narayana, N., Shamala, N., Ganesh, K.N., Viswamitra, M.A.(2006) Biochemistry 45: 1200-1211
- PubMed: 16430216 Search on PubMed
- DOI: https://doi.org/10.1021/bi051569l
- Primary Citation Related Structures:
2F8W - PubMed Abstract:
The crystal structure of a hexamer duplex d(CACGTG)(2) has been determined and refined to an R-factor of 18.3% using X-ray data up to 1.2 A resolution. The sequence crystallizes as a left-handed Z-form double helix with Watson-Crick base pairing. There is one hexamer duplex, a spermine molecule, 71 water molecules, and an unexpected diamine (Z-5, 1,3-propanediamine, C(3)H(10)N(2)) in the asymmetric unit. This is the high-resolution non-disordered structure of a Z-DNA hexamer containing two AT base pairs in the interior of a duplex with no modifications such as bromination or methylation on cytosine bases. This structure does not possess multivalent cations such as cobalt hexaammine that are known to stabilize Z-DNA. The overall duplex structure and its crystal interactions are similar to those of the pure-spermine form of the d(CGCGCG)(2) structure. The spine of hydration in the minor groove is intact except in the vicinity of the T5A8 base pair. The binding of the Z-5 molecule in the minor grove of the d(CACGTG)(2) duplex appears to have a profound effect in conferring stability to a Z-DNA conformation via electrostatic complementarity and hydrogen bonding interactions. The successive base stacking geometry in d(CACGTG)(2) is similar to the corresponding steps in d(CG)(3). These results suggest that specific polyamines such as Z-5 could serve as powerful inducers of Z-type conformation in unmodified DNA sequences with AT base pairs. This structure provides a molecular basis for stabilizing AT base pairs incorporated into an alternating d(CG) sequence.
- Department of Biochemistry, Case Western Reserve University, Cleveland, Ohio 44106, USA. nxn17@case.edu
Organizational Affiliation: