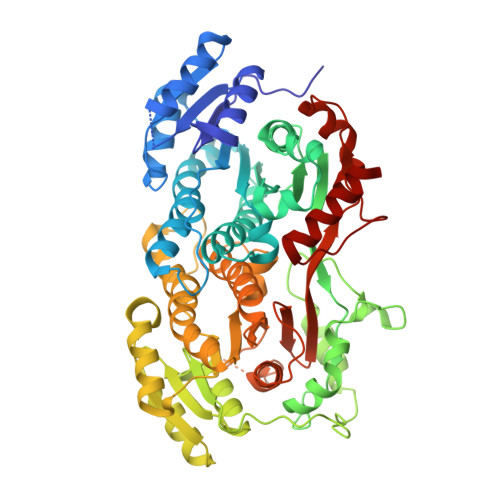
Crystal Structure of Yeast Peroxisomal Multifunctional Enzyme: Structural Basis for Substrate Specificity of (3R)-hydroxyacyl-CoA Dehydrogenase Units.
Ylianttila, M.S., Pursiainen, N.V., Haapalainen, A.M., Juffer, A.H., Poirier, Y., Hiltunen, J.K., Glumoff, T.(2006) J Mol Biology 358: 1286-1295
- PubMed: 16574148 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.03.001
- Primary Citation Related Structures:
2ET6 - PubMed Abstract:
(3R)-hydroxyacyl-CoA dehydrogenase is part of multifunctional enzyme type 2 (MFE-2) of peroxisomal fatty acid beta-oxidation. The MFE-2 protein from yeasts contains in the same polypeptide chain two dehydrogenases (A and B), which possess difference in substrate specificity. The crystal structure of Candida tropicalis (3R)-hydroxyacyl-CoA dehydrogenase AB heterodimer, consisting of dehydrogenase A and B, determined at the resolution of 2.2A, shows overall similarity with the prototypic counterpart from rat, but also important differences that explain the substrate specificity differences observed. Docking studies suggest that dehydrogenase A binds the hydrophobic fatty acyl chain of a medium-chain-length ((3R)-OH-C10) substrate as bent into the binding pocket, whereas the short-chain substrates are dislocated by two mechanisms: (i) a short-chain-length 3-hydroxyacyl group ((3R)-OH-C4) does not reach the hydrophobic contacts needed for anchoring the substrate into the active site; and (ii) Leu44 in the loop above the NAD(+) cofactor attracts short-chain-length substrates away from the active site. Dehydrogenase B, which can use a (3R)-OH-C4 substrate, has a more shallow binding pocket and the substrate is correctly placed for catalysis. Based on the current structure, and together with the structure of the 2-enoyl-CoA hydratase 2 unit of yeast MFE-2 it becomes obvious that in yeast and mammalian MFE-2s, despite basically identical functional domains, the assembly of these domains into a mature, dimeric multifunctional enzyme is very different.
- Biocenter Oulu and Department of Biochemistry, University of Oulu, PO Box 3000, FIN-90014, University of Oulu, Finland.
Organizational Affiliation: