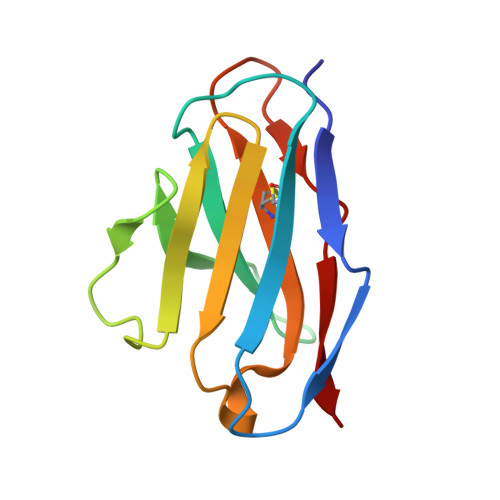
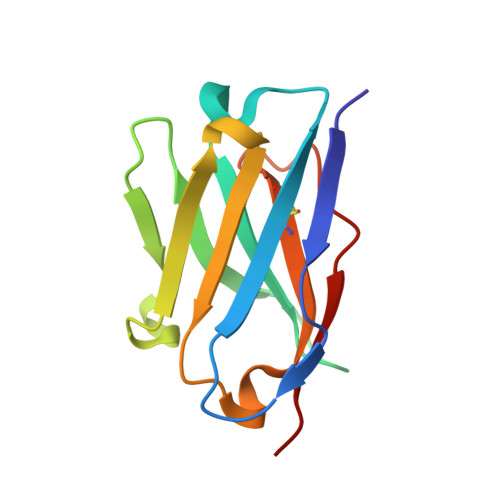
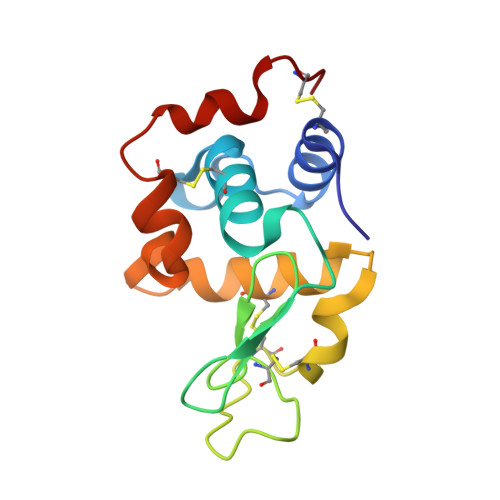
Critical contribution of VH-VL interaction to reshaping of an antibody: the case of humanization of anti-lysozyme antibody, HyHEL-10
Nakanishi, T., Tsumoto, K., Yokota, A., Kondo, H., Kumagai, I.(2008) Protein Sci 17: 261-270
- PubMed: 18227432 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.073156708
- Primary Citation Related Structures:
2EIZ, 2EKS, 2YSS - PubMed Abstract:
To clarify the effects of humanizing a murine antibody on its specificity and affinity for its target, we examined the interaction between hen egg white lysozyme (HEL) and its antibody, HyHEL-10 variable domain fragment (Fv). We selected a human antibody framework sequence with high homology, grafted sequences of six complementarity-determining regions of murine HyHEL-10 onto the framework, and investigated the interactions between the mutant Fvs and HEL. Isothermal titration calorimetry indicated that the humanization led to 10-fold reduced affinity of the antibody for its target, due to an unfavorable entropy change. Two mutations together into the interface of the variable domains, however, led to complete recovery of antibody affinity and specificity for the target, due to reduction of the unfavorable entropy change. X-ray crystallography of the complex of humanized antibodies, including two mutants, with HEL demonstrated that the complexes had almost identical structures and also paratope and epitope residues were almost conserved, except for complementary association of variable domains. We conclude that adjustment of the interfacial structures of variable domains can contribute to the reversal of losses of affinity or specificity caused by humanization of murine antibodies, suggesting that appropriate association of variable domains is critical for humanization of murine antibodies without loss of function.
- Department of Biomolecular Engineering, Graduate School of Engineering, Tohoku University, Sendai 980-8579, Japan.
Organizational Affiliation: