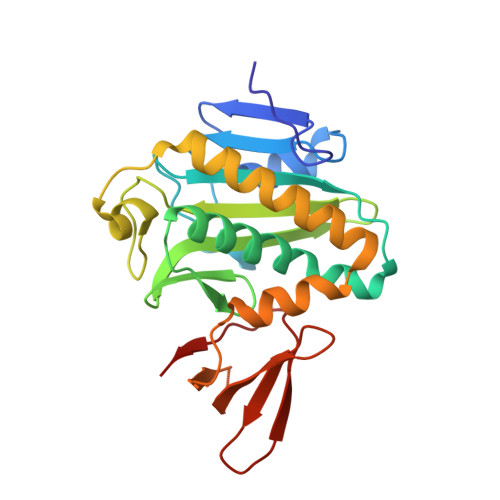
Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
Bagautdinov, B., Matsuura, Y., Bagautdinova, S., Kunishima, N.(2008) J Biological Chem 283: 14739-14750
- PubMed: 18372281 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M709116200
- Primary Citation Related Structures:
1X01, 2D5D, 2DXU, 2DZC, 2E41, 2E64, 2EJF, 2EJG, 2EVB, 2ZGW - PubMed Abstract:
Biotin protein ligase (BPL) catalyzes the biotinylation of the biotin carboxyl carrier protein (BCCP) only at a special lysine residue. Here we report the first structure of BPL.BCCP complex crystals, which are prepared using two BPL mutants: R48A and R48A/K111A. From a detailed structural characterization, it is likely that the mutants retain functionality as enzymes but have a reduced activity to produce the reaction intermediate biotinyl-5'-AMP. The observed biotin and partly disordered ATP in the mutant structures may act as a non-reactive analog of the substrates or biotinyl-5'-AMP, thereby providing the complex crystals. The four crystallographically independent BPL.BCCP complexes obtained can be classified structurally into three groups: the formation stages 1 and 2 with apo-BCCP and the product stage with biotinylated holo-BCCP. Residues responsible for the complex formation as well as for the biotinylation reaction have been identified. The C-terminal domain of BPL shows especially large conformational changes to accommodate BCCP, suggesting its functional importance. The formation stage 1 complex shows the closest distance between the carboxyl carbon of biotin and the special lysine of BCCP, suggesting its relevance to the unobserved reaction stage. Interestingly, bound ATP and biotin are also seen in the product stage, indicating that the substrates may be recruited into the product stage complex before the release of holo-BCCP, probably for the next reaction cycle. The existence of formation and product stages before and after the reaction stage would be favorable to ensure both the reaction efficiency and the extreme substrate specificity of the biotinylation reaction.
- RIKEN SPring-8 Center, Harima Institute, Sayo-Gun, Hyogo, Japan.
Organizational Affiliation: