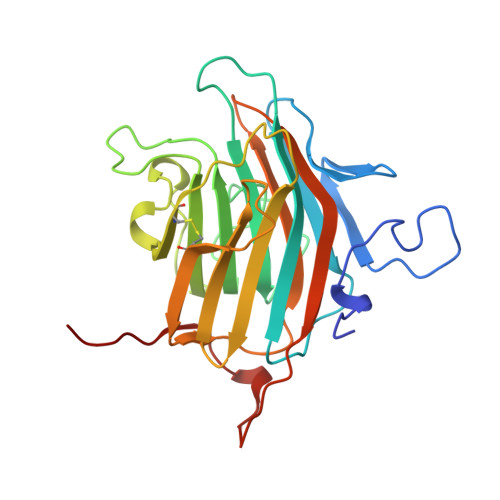
Structural basis for recognition of high mannose type glycoproteins by mammalian transport lectin VIP36
Satoh, T., Cowieson, N.P., Hakamata, W., Ideo, H., Fukushima, K., Kurihara, M., Kato, R., Yamashita, K., Wakatsuki, S.(2007) J Biological Chem 282: 28246-28255
- PubMed: 17652092 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M703064200
- Primary Citation Related Structures:
2DUO, 2DUP, 2DUQ, 2DUR, 2E6V - PubMed Abstract:
VIP36 functions as a transport lectin for trafficking certain high mannose type glycoproteins in the secretory pathway. Here we report the crystal structure of VIP36 exoplasmic/luminal domain comprising a carbohydrate recognition domain and a stalk domain. The structures of VIP36 in complex with Ca(2+) and mannosyl ligands are also described. The carbohydrate recognition domain is composed of a 17-stranded antiparallel beta-sandwich and binds one Ca(2+) adjoining the carbohydrate-binding site. The structure reveals that a coordinated Ca(2+) ion orients the side chains of Asp(131), Asn(166), and His(190) for carbohydrate binding. This result explains the Ca(2+)-dependent carbohydrate binding of this protein. The Man-alpha-1,2-Man-alpha-1,2-Man, which corresponds to the D1 arm of high mannose type glycan, is recognized by eight residues through extensive hydrogen bonds. The complex structures reveal the structural basis for high mannose type glycoprotein recognition by VIP36 in a Ca(2+)-dependent and D1 arm-specific manner.
- Structural Biology Research Center, Photon Factory, Institute of Materials Structure Science, High Energy Accelerator Research Organization (KEK), Tsukuba, Ibaraki 305-0801, Japan.
Organizational Affiliation: