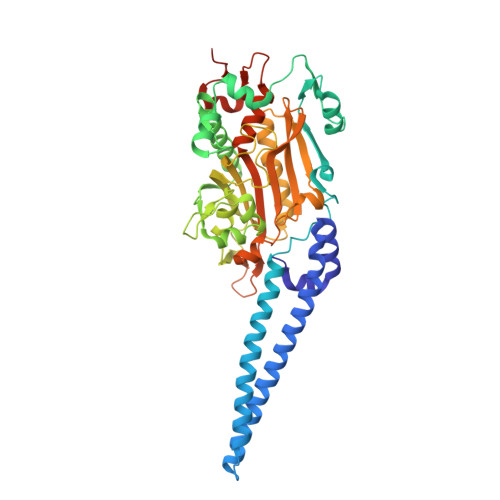
Crystallographic and mutational studies of seryl-tRNA synthetase from the archaeon Pyrococcus horikoshii.
Itoh, Y., Sekine, S., Kuroishi, C., Terada, T., Shirouzu, M., Kuramitsu, S., Yokoyama, S.(2008) RNA Biol 5: 169-177
- PubMed: 18818520 Search on PubMed
- DOI: https://doi.org/10.4161/rna.5.3.6876
- Primary Citation Related Structures:
2DQ0, 2ZR2, 2ZR3 - PubMed Abstract:
Seryl-tRNA synthetase (SerRS) catalyzes the ligation of serine to the 3'-end of serine tRNA (tRNA(Ser)), which is typical of the type-2 tRNAs characterized by a long extra arm. The SerRSs are divided into two types, the archaeal/eukaryal and bacterial types. In this study, we solved the crystal structures of the SerRS from the archaeon Pyrococcus horikoshii bound with 5'-O-[N-(L-seryl)-sulfamoyl]-adenosine at 2.6 A and with ATP at 2.8 A, as well as in the apo form at 3.0 A. P. horikoshii SerRS recognizes the seryl and adenylate moieties in a manner similar to those of the bacterial and mitochondrial SerRSs from Thermus thermophilus and Bos taurus, respectively, but different from that of the unusual SerRS from the methanogenic archaeon Methanosarcina barkeri. P. horikoshii SerRS efficiently aminoacylated not only P. horikoshii tRNA(Ser) but also bacterial tRNA(Ser)s from T. thermophilus and Escherichia coli. Models of P. horikoshii SerRS bound with the T. thermophilus and P. horikoshii tRNA(Ser)s suggested that the helical domain of P. horikoshii SerRS is involved in the extra arm binding. This region of P. horikoshii SerRS has additional basic residues as compared with T. thermophilus SerRS, and a Trp residue specific to the archaeal/eukaryal SerRSs. Mutational analyses revealed that the basic and Trp residues are important for tRNA aminoacylation. P. horikoshii SerRS has the archaea-specific insertion, which collaborates with the core domain to form a basic channel leading to the active site. Two sulfate ions are bound to the channel, suggesting that the tRNA 3' region might bind to the channel.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: