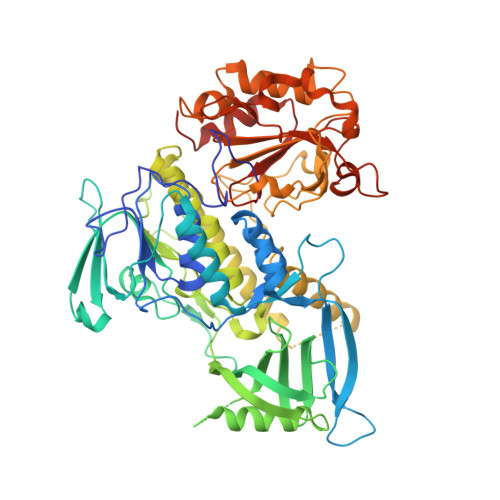
Crystal structure of 3-hydroxybenzoate hydroxylase from Comamonas testosteroni has a large tunnel for substrate and oxygen access to the active site
Hiromoto, T., Fujiwara, S., Hosokawa, K., Yamaguchi, H.(2006) J Mol Biology 364: 878-896
- PubMed: 17045293 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.09.031
- Primary Citation Related Structures:
2DKH, 2DKI - PubMed Abstract:
The 3-hydroxybenzoate hydroxylase (MHBH) from Comamonas testosteroni KH122-3s is a single-component flavoprotein monooxygenase, a member of the glutathione reductase (GR) family. It catalyzes the conversion of 3-hydroxybenzoate to 3,4-dihydroxybenzoate with concomitant requirements for equimolar amounts of NADPH and molecular oxygen. The production of dihydroxy-benzenoid derivative by hydroxylation is the first step in the aerobic degradation of various phenolic compounds in soil microorganisms. To establish the structural basis for substrate recognition, the crystal structure of MHBH in complex with its substrate was determined at 1.8 A resolution. The enzyme is shown to form a physiologically active homodimer with crystallographic 2-fold symmetry, in which each subunit consists of the first two domains comprising an active site and the C-terminal domain involved in oligomerization. The protein fold of the catalytic domains and the active-site architecture, including the FAD and substrate-binding sites, are similar to those of 4-hydroxybenzoate hydroxylase (PHBH) and phenol hydroxylase (PHHY), which are members of the GR family, providing evidence that the flavoprotein aromatic hydroxylases share similar catalytic actions for hydroxylation of the respective substrates. Structural comparison of MHBH with the homologous enzymes suggested that a large tunnel connecting the substrate-binding pocket to the protein surface serves for substrate transport in this enzyme. The internal space of the large tunnel is distinctly divided into hydrophilic and hydrophobic regions. The characteristically stratified environment in the tunnel interior and the size of the entrance would allow the enzyme to select its substrate by amphiphilic nature and molecular size. In addition, the structure of the Xe-derivative at 2.5 A resolution led to the identification of a putative oxygen-binding site adjacent to the substrate-binding pocket. The hydrophobic nature of the xenon-binding site extends to the solvent through the tunnel, suggesting that the tunnel could be involved in oxygen transport.
- Department of Chemistry, Nanobiotechnology Research Center, School of Science and Technology, Kwansei Gakuin University, 2-1 Gakuen, Sanda, Hyogo 669-1337, Japan.
Organizational Affiliation: