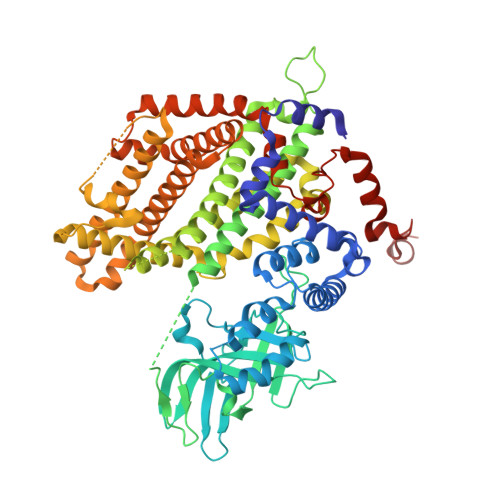
Three-Dimensional Structure of Rat-Liver Acyl-CoA Oxidase in Complex with a Fatty Acid: Insights into Substrate-Recognition and Reactivity toward Molecular Oxygen.
Tokuoka, K., Nakajima, Y., Hirotsu, K., Miyahara, I., Nishina, Y., Shiga, K., Tamaoki, H., Setoyama, C., Tojo, H., Miura, R.(2006) J Biochem 139: 789-795
- PubMed: 16672280 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvj088
- Primary Citation Related Structures:
2DDH - PubMed Abstract:
The three-dimensional structure of rat-liver acyl-CoA oxidase-II (ACO-II) in a complex with a C12-fatty acid was solved by the molecular replacement method based on the uncomplexed ACO-II structure. The crystalline form of the complex was obtained by cocrystallization of ACO-II with dodecanoyl-CoA. The crystalline complex possessed, in the active-site crevice, only the fatty acid moiety that had been formed through hydrolysis of the thioester bond. The overall dimeric structure and the folding pattern of each subunit are essentially superimposable on those of uncomplexed ACO-II. The active site including the flavin ring of FAD, the crevice embracing the fatty acyl moiety, and adjacent amino acid side chains are superimposably conserved with the exception of Glu421, whose carboxylate group is tilted away to accommodate the fatty acid. One of the carboxyl oxygens of the bound fatty acid is hydrogen-bonded to the amide hydrogen of Glu421, the presumed catalytic base, and to the ribityl 2'-hydroxyl group of FAD. This hydrogen-bonding network correlates well with the substrate recognition/activation in acyl-CoA dehydrogenase. The binding mode of C12-fatty acid suggests that the active site does not close upon substrate binding, but remains spacious during the entire catalytic process, the oxygen accessibility in the oxidative half-reaction thereby being maintained.
- Department of Chemistry, Graduate School of Science, Osaka City University, Sugimoto, Sumiyoshi-ku, Osaka 558-8585.
Organizational Affiliation: