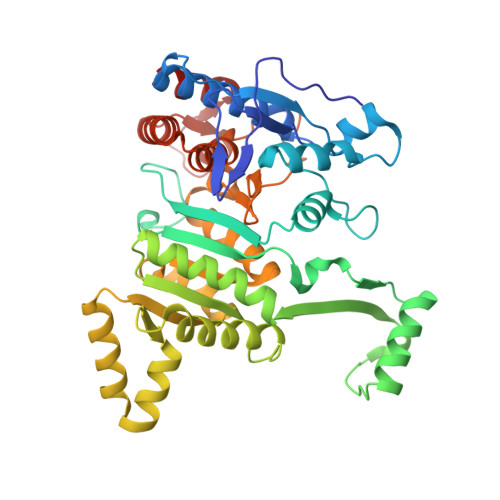
Structure and quantum chemical analysis of NAD+-dependent isocitrate dehydrogenase: hydride transfer and co-factor specificity
Imada, K., Tamura, T., Takenaka, R., Kobayashi, I., Namba, K., Inagaki, K.(2008) Proteins 70: 63-71
- PubMed: 17634983
- DOI: https://doi.org/10.1002/prot.21486
- Primary Citation Related Structures:
2D4V - PubMed Abstract:
The crystal structure of Acidithiobacillus thiooxidans isocitrate dehydrogenase complexed with NAD+ and citrate has been solved to a resolution of 1.9 A. The protein fold of this NAD+-dependent enzyme shares a high similarity with those of NADP+-dependent bacterial ICDHs. The NAD+ and the citrate are clearly identified in the active site cleft with a well-defined electron density. Asp-357 is the direct cofactor-specificity determinant that interacts with 2'-OH and 3'-OH of the adenosine ribose. The adenosine ribose takes a C2'-endo puckering conformation as previously reported for an NAD+-specific isopropylmalate dehydrogenase. The nicotinamide moiety of NAD+ has the amide NH2 group oriented in cis conformation with respect to the C4 carbon of the nicotinamide ring, slanted toward the bound citrate molecule with a dihedral angle of -21 degrees . The semi-empirical molecular orbital calculation suggests that the pro-R hydrogen atom at C4 of NADH would bear the largest negative charge when the amide NH2 group is in such conformation, suggesting that the amide group has a catalytically significant role in stabilizing the transition state as NADH is being formed during the hydride transfer catalysis.
- Department of Frontier Biosciences, Osaka University, 1-3 Yamadaoka, Suita, Osaka 565-0871, Japan. kimada@fbs.osaka-u.ac.jp
Organizational Affiliation: