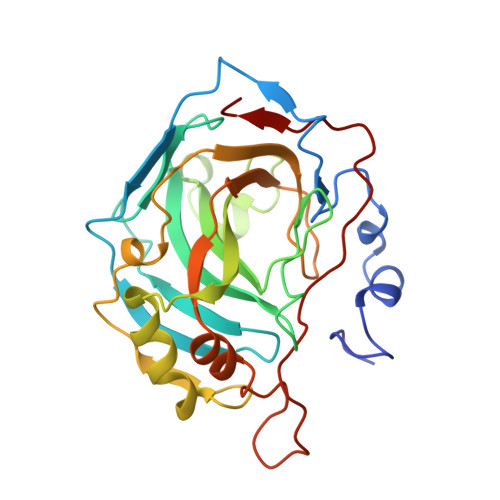
Structure of native and apo carbonic anhydrase II and structure of some of its anion-ligand complexes.
Hakansson, K., Carlsson, M., Svensson, L.A., Liljas, A.(1992) J Mol Biology 227: 1192-1204
- PubMed: 1433293 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90531-n
- Primary Citation Related Structures:
2CBA, 2CBB, 2CBC, 2CBD, 2CBE - PubMed Abstract:
In order to obtain a better structural framework for understanding the catalytic mechanism of carbonic anhydrase, a number of inhibitor complexes of the enzyme were investigated crystallographically. The three-dimensional structure of free human carbonic anhydrase II was refined at pH 7.8 (1.54 A resolution) and at pH 6.0 (1.67 A resolution). The structure around the zinc ion was identical at both pH values. The structure of the zinc-free enzyme was virtually identical with that of the native enzyme, apart from a water molecule that had moved 0.9 A to fill the space that would be occupied by the zinc ion. The complexes with the anionic inhibitors bisulfite and formate were also studied at neutral pH. Bisulfite binds with one of its oxygen atoms, presumably protonized, to the zinc ion and replaces the zinc water. Formate, lacking a hydroxyl group, is bound with its oxygen atoms not far away from the position of the non-protonized oxygen atoms of the bisulfite complex, i.e. at hydrogen bond distance from Thr199 N and at a position between the zinc ion and the hydrophobic part of the active site. The result of these and other studies have implications for our view of the catalytic function of the enzyme, since virtually all inhibitors share some features with substrate, product or expected transition states. A reaction scheme where electrophilic activation of carbon dioxide plays an important role in the hydration reaction is presented. In the reverse direction, the protonized oxygen of the bicarbonate is forced upon the zinc ion, thereby facilitating cleavage of the carbon-oxygen bond. This is achieved by the combined action of the anionic binding site, which binds carboxyl groups, the side-chain of threonine 199, which discriminates between hydrogen bond donors and acceptors, and hydrophobic interaction between substrate and the active site cavity. The required proton transfer between the zinc water and His64 can take place through water molecules 292 and 318.
- Chemical Center, University of Lund, Sweden.
Organizational Affiliation: