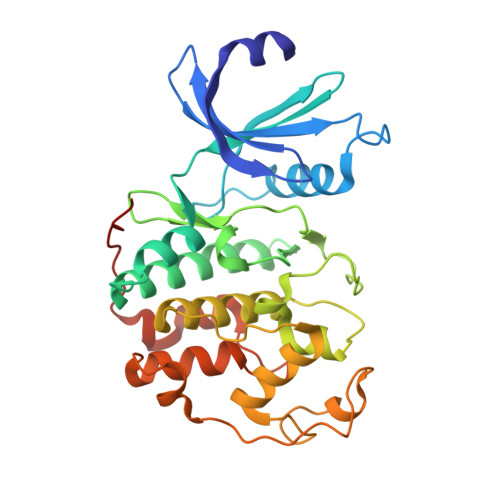
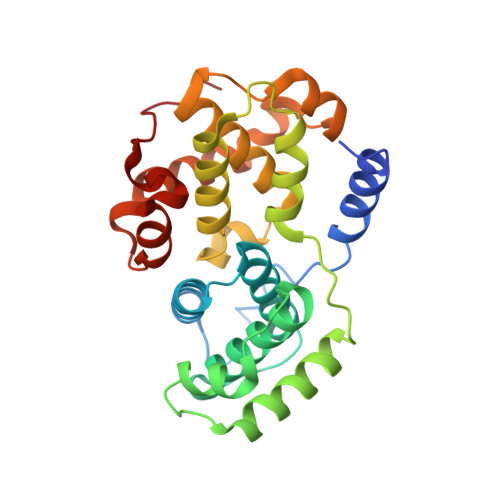
3-Amino-1,4,5,6-Tetrahydropyrrolo[3,4-C]Pyrazoles: A New Class of Cdk2 Inhibitors.
Pevarello, P., Fancelli, D., Vulpetti, A., Amici, R., Villa, M., Pittala, V., Vianello, P., Cameron, A., Ciomei, M., Mercurio, C., Bischoff, J.R., Roletto, F., Varasi, M., Brasca, M.G.(2006) Bioorg Med Chem Lett 16: 1084
- PubMed: 16290148 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2005.10.071
- Primary Citation Related Structures:
2C4G - PubMed Abstract:
We have recently reported about a new class of Aurora-A inhibitors based on a bicyclic tetrahydropyrrolo[3,4-c]pyrazole scaffold. Here we describe the synthesis and early expansion of CDK2/cyclin A-E inhibitors belonging to the same chemical class. Synthesis of the compounds was accomplished using a solution-phase protocol amenable to rapid parallel expansion. Compounds with nanomolar activity in the biochemical assay and able to efficiently inhibit CDK2-mediated tumor cell proliferation have been obtained.
- Department of Chemistry, Nerviano Medical Science, BU-Oncology, Via Pasteur 10, 20014 Nerviano MI, Italy. paolo.pevarello@nervianoms.com
Organizational Affiliation: