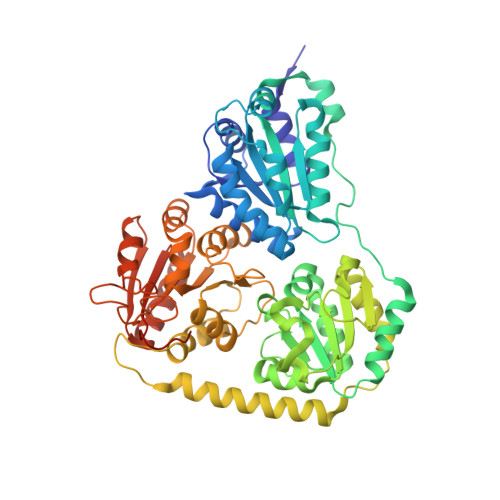
Structural Basis for Activation of the Thiamin Diphosphate-Dependent Enzyme Oxalyl-Coa Decarboxylase by Adenosine Diphosphate.
Berthold, C.L., Moussatche, P., Richards, N.G.J., Lindqvist, Y.(2005) J Biological Chem 280: 41645
- PubMed: 16216870
- DOI: https://doi.org/10.1074/jbc.M509921200
- Primary Citation Related Structures:
2C31 - PubMed Abstract:
Oxalyl-coenzyme A decarboxylase is a thiamin diphosphate-dependent enzyme that plays an important role in the catabolism of the highly toxic compound oxalate. We have determined the crystal structure of the enzyme from Oxalobacter formigenes from a hemihedrally twinned crystal to 1.73 A resolution and characterized the steady-state kinetic behavior of the decarboxylase. The monomer of the tetrameric enzyme consists of three alpha/beta-type domains, commonly seen in this class of enzymes, and the thiamin diphosphate-binding site is located at the expected subunit-subunit interface between two of the domains with the cofactor bound in the conserved V-conformation. Although oxalyl-CoA decarboxylase is structurally homologous to acetohydroxyacid synthase, a molecule of ADP is bound in a region that is cognate to the FAD-binding site observed in acetohydroxyacid synthase and presumably fulfils a similar role in stabilizing the protein structure. This difference between the two enzymes may have physiological importance since oxalyl-CoA decarboxylation is an essential step in ATP generation in O. formigenes, and the decarboxylase activity is stimulated by exogenous ADP. Despite the significant degree of structural conservation between the two homologous enzymes and the similarity in catalytic mechanism to other thiamin diphosphate-dependent enzymes, the active site residues of oxalyl-CoA decarboxylase are unique. A suggestion for the reaction mechanism of the enzyme is presented.
- Molecular Structural Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17177 Stockholm, Sweden.
Organizational Affiliation: