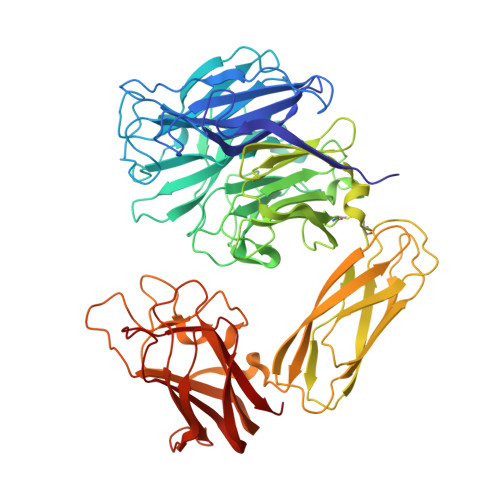
Galactose Recognition by the Carbohydrate-Binding Module of a Bacterial Sialidase.
Newstead, S.L., Watson, J.N., Bennet, A.J., Taylor, G.(2005) Acta Crystallogr D Biol Crystallogr 61: 1483
- PubMed: 16239725 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905026132
- Primary Citation Related Structures:
2BZD - PubMed Abstract:
Glycoside hydrolases often possess carbohydrate-binding modules (CBMs) in addition to their catalytic domains, which help target the enzymes to appropriate substrates and thereby increase their catalytic efficiency. Sialidases hydrolyse the release of sialic acid from a variety of glycoconjugates and play significant roles in the pathogenesis of a number of important diseases. The sialidase from Micromonospora viridifaciens has a CBM which recognizes galactose. The CBM is linked to the catalytic domain by an immunoglobulin-like domain, resulting in the galactose binding site sitting above the catalytic site, suggesting an interplay between the two sites. By studying nine crystallographically independent structures of the M. viridifaciens sialidase, the relative flexibility of the three domains was analysed. A detailed study is also presented of the recognition of galactose and lactose by the M. viridifaciens CBM. The striking structure of this sialidase suggests a role for the CBM in binding to galactose residues unmasked by the adjacent catalytic site.
- Centre for Biomolecular Sciences, University of St Andrews, St Andrews, Fife KY16 9ST, Scotland.
Organizational Affiliation: