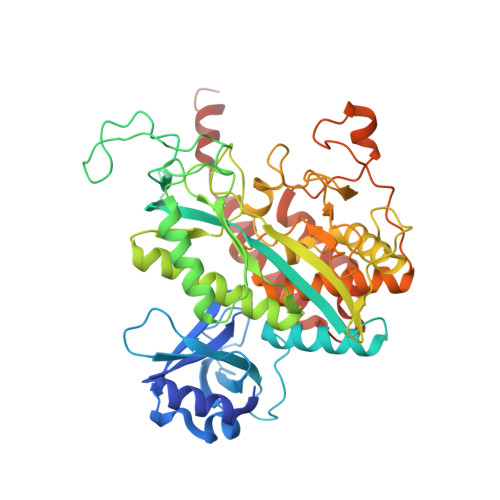
Structure of Mycobacterium Tuberculosis Glutamine Synthetase in Complex with a Transition-State Mimic Provides Functional Insights.
Krajewski, W.W., Jones, T.A., Mowbray, S.L.(2005) Proc Natl Acad Sci U S A 102: 10499
- PubMed: 16027359 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0502248102
- Primary Citation Related Structures:
2BVC - PubMed Abstract:
Glutamine synthetase catalyzes the ligation of glutamate and ammonia to form glutamine, with the resulting hydrolysis of ATP. The enzyme is a central component of bacterial nitrogen metabolism and is a potential drug target. Here, we report a high-yield recombinant expression system for glutamine synthetase of Mycobacterium tuberculosis together with a simple purification. The procedure allowed the structure of a complex with a phosphorylated form of the inhibitor methionine sulfoximine, magnesium, and ADP to be solved by molecular replacement and refined at 2.1-A resolution. To our knowledge, this study provides the first reported structure for a taut form of the M. tuberculosis enzyme, similar to that observed for the Salmonella enzyme earlier. The phospho compound, generated in situ by an active enzyme, mimics the phosphorylated tetrahedral adduct at the transition state. Some differences in ligand interactions of the protein with both phosphorylated compound and nucleotide are observed compared with earlier structures; a third metal ion also is found. The importance of these differences in the catalytic mechanism is discussed; the results will help guide the search for specific inhibitors of potential therapeutic interest.
- Department of Cell and Molecular Biology, Uppsala University, Biomedical Center, Box 596, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: