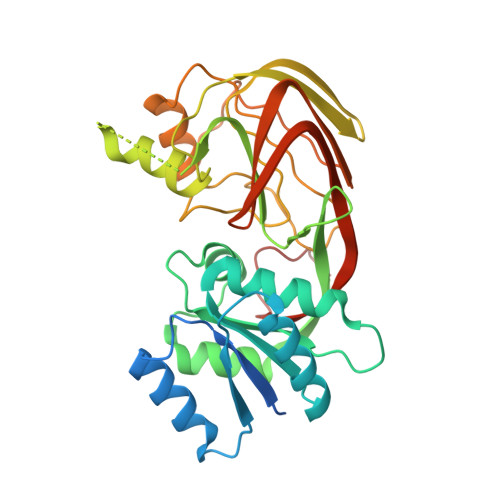
Crystal Structure of Yegs, a Homologue to the Mammalian Diacylglycerol Kinases, Reveals a Novel Regulatory Metal Binding Site.
Bakali, H.M., Herman, M.D., Johnson, K.A., Kelly, A.A., Wieslander, A., Hallberg, B.M., Nordlund, P.(2007) J Biological Chem 282: 19644
- PubMed: 17351295 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M604852200
- Primary Citation Related Structures:
2BON, 2JGR - PubMed Abstract:
The human lipid kinase family controls cell proliferation, differentiation, and tumorigenesis and includes diacylglycerol kinases, sphingosine kinases, and ceramide kinases. YegS is an Escherichia coli protein with significant sequence homology to the catalytic domain of the human lipid kinases. We have solved the crystal structure of YegS and shown that it is a lipid kinase with phosphatidylglycerol kinase activity. The crystal structure reveals a two-domain protein with significant structural similarity to a family of NAD kinases. The active site is located in the interdomain cleft formed by four conserved sequence motifs. Surprisingly, the structure reveals a novel metal binding site composed of residues conserved in most lipid kinases.
- Division of Biophysics, Department of Medical Biochemistry and Biophysics, Karolinska Institute, SE-171 77 Stockholm, Sweden.
Organizational Affiliation: