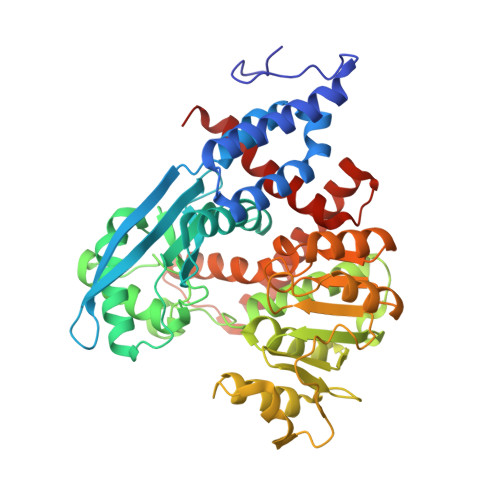
The Crystal Structure of Plasmodium Falciparum Glutamate Dehydrogenase, a Putative Target for Novel Antimalarial Drugs
Werner, C., Stubbs, M.T., Krauth-Siege, R.L., Klebe, G.(2005) J Mol Biology 349: 597
- PubMed: 15878595 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.03.077
- Primary Citation Related Structures:
2BMA - PubMed Abstract:
Plasmodium falciparum is the main causative agent of tropical malaria, the most severe parasitic disease in the world. Growing resistance of Plasmodia towards available drugs is an increasing problem in countries where malaria is endemic. As Plasmodia are sensitive to oxidative stress, augmenting this in the parasite represents a promising principle for the development of novel antimalarial drugs. The NADP-dependent glutamate dehydrogenase (GDH) of P.falciparum is largely responsible for the production of NADPH in the parasite, which in turn serves as electron source for the antioxidative enzymes glutathione reductase and thioredoxin reductase. As GDH does not occur in the host erythrocyte, GDH is a particularly attractive target for drug therapy. The three-dimensional structure of P.falciparum GDH in the unligated state has been determined by X-ray crystallography to a resolution of 2.7A. Compared to the mammalian enzymes, two amino acid residues are exchanged in the putative active site of the parasite GDH. The most obvious differences between parasite and human GDH are the subunit interfaces of the hexameric proteins. In the parasite protein, several salt-bridges mediate contacts between the subunits whereas in the human enzyme these interactions are mainly of hydrophobic nature. Furthermore, P.falciparum GDH possesses a unique N-terminal extension that does not occur in any other GDH sequence so far studied. These findings might be exploited for the design of peptidomimetics capable of disrupting the oligomeric organisation of the parasite enzyme.
- Institute of Pharmaceutical Chemistry, Philipps-University Marburg, Marbacher Weg 6, D-35037 Marburg, Germany.
Organizational Affiliation: