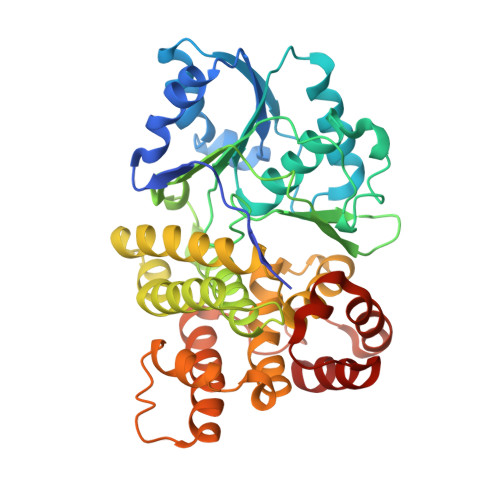
Crystal Structure of an Iron-Dependent Group III Dehydrogenase that Interconverts L-Lactaldehyde and L-1,2-Propanediol in Escherichia Coli
Montella, C., Bellsolell, L., Perez-Luque, R., Badia, J., Baldoma, L., Coll, M., Aguilar, J.(2005) J Bacteriol 187: 4957
- PubMed: 15995211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.187.14.4957-4966.2005
- Primary Citation Related Structures:
2BI4, 2BL4 - PubMed Abstract:
The FucO protein, a member of the group III "iron-activated" dehydrogenases, catalyzes the interconversion between L-lactaldehyde and L-1,2-propanediol in Escherichia coli. The three-dimensional structure of FucO in a complex with NAD(+) was solved, and the presence of iron in the crystals was confirmed by X-ray fluorescence. The FucO structure presented here is the first structure for a member of the group III bacterial dehydrogenases shown experimentally to contain iron. FucO forms a dimer, in which each monomer folds into an alpha/beta dinucleotide-binding N-terminal domain and an all-alpha-helix C-terminal domain that are separated by a deep cleft. The dimer is formed by the swapping (between monomers) of the first chain of the beta-sheet. The binding site for Fe(2+) is located at the face of the cleft formed by the C-terminal domain, where the metal ion is tetrahedrally coordinated by three histidine residues (His200, His263, and His277) and an aspartate residue (Asp196). The glycine-rich turn formed by residues 96 to 98 and the following alpha-helix is part of the NAD(+) recognition locus common in dehydrogenases. Site-directed mutagenesis and enzyme kinetic assays were performed to assess the role of different residues in metal, cofactor, and substrate binding. In contrast to previous assumptions, the essential His267 residue does not interact with the metal ion. Asp39 appears to be the key residue for discriminating against NADP(+). Modeling L-1,2-propanediol in the active center resulted in a close approach of the C-1 hydroxyl of the substrate to C-4 of the nicotinamide ring, implying that there is a typical metal-dependent dehydrogenation catalytic mechanism.
- Department of Biochemistry, School of Pharmacy, University of Barcelona, Spain.
Organizational Affiliation: