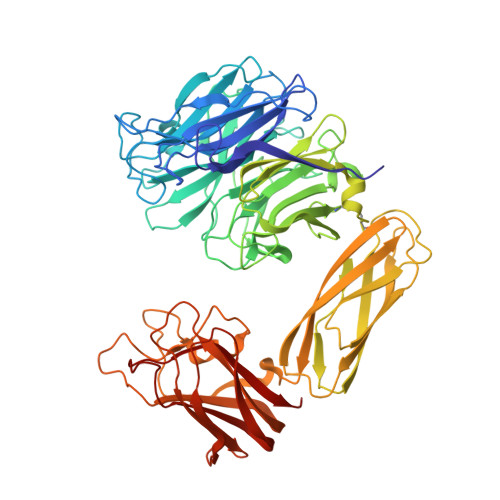
Structure and Mechanism of Action of an Inverting Mutant Sialidase.
Newstead, S., Watson, J.N., Knoll, T.L., Bennet, A.J., Taylor, G.L.(2005) Biochemistry 44: 9117
- PubMed: 15966735 Search on PubMed
- DOI: https://doi.org/10.1021/bi050517t
- Primary Citation Related Structures:
2BER - PubMed Abstract:
Mutagenesis of the conserved tyrosine (Y370) of the Micromonospora viridifaciens sialidase to small amino acids changes the mechanism of catalysis from retention of anomeric configuration to inversion [Watson, J. N., et al. (2003) Biochemistry 42, 12682-12690]. For the Y370G mutant enzyme-catalyzed hydrolysis of a series of aryl sialosides and 3'-sialyllactose, the derived Brønsted parameters (beta(lg)) on k(cat) and k(cat)/K(m) are -0.63 +/- 0.05 and -0.80 +/- 0.08, respectively. Thus, for the Y370G enzyme, glycosidic C-O bond cleavage is rate-determining. Analysis of the activity of the Y370G mutant and wild-type enzymes against a substrate [3,4-dihydro-2H-pyrano[3,2-c]pyridinium alpha-d-N-acetylneuraminide (DHP-alphaNeu5Ac)] whose hydrolysis cannot be accelerated by acid catalysis is consistent with these reactions proceeding via S(N)1 and S(N)2 mechanisms, respectively. The overall structure of the Y370G mutant sialidase active site is very similar to the previously reported wild-type structure [Gaskell, A., et al. (1995) Structure 3, 1197-1205], although removal of the tyrosine residue creates two significant changes to the active site. First, the anomeric oxygen atom of the hydrolysis product (beta-N-acetylneuraminic acid) and four water molecules bind in the large cavity created by the Y370G mutation. Second, the side chain of Asn310 moves to make a strong hydrogen bond to one of the bound water molecules.
- Centre for Biomolecular Science, University of St. Andrews, St. Andrews, Fife KY16 9ST, Scotland.
Organizational Affiliation: