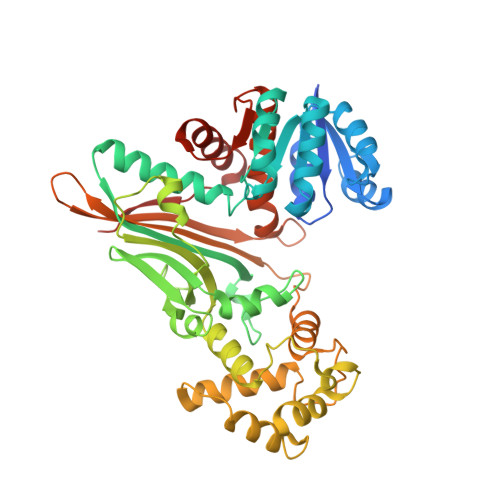
Crystal structure of the his-tagged saccharopine reductase from Saccharomyces cerevisiae at 1.7-A resolution.
Andi, B., Cook, P.F., West, A.H.(2006) Cell Biochem Biophys 46: 17-26
- PubMed: 16943620 Search on PubMed
- DOI: https://doi.org/10.1385/CBB:46:1:17
- Primary Citation Related Structures:
2AXQ - PubMed Abstract:
The three-dimensional structure of the saccharopine reductase enzyme from the budding yeast Saccharomyces cerevisiae was determined to 1.7-A resolution in the apo form by using molecular replacement. The enzyme monomer consists of three domains: domain I is a variant of the Rossmann fold, domain II folds into a alpha/beta structure containing a mixed seven-stranded beta-sheet as the central core, and domain III has an all-helical fold. Comparative fold alignment with the enzyme from Magnaporthe grisea suggests that domain I binds to NADPH, and domain II binds to saccharopine and is involved in dimer formation. Domain III is involved in closing the active site of the enzyme once substrates are bound. Structural comparison of the saccharopine reductase enzymes from S. cerevisiae and M. grisea indicates that domain II has the highest number of conserved residues, suggesting that it plays an important role in substrate binding and in spatially orienting domains I and III.
- Department of Chemistry and Biochemistry, University of Oklahoma, 620 Parrington Oval, Norman, OK 73019, USA.
Organizational Affiliation: