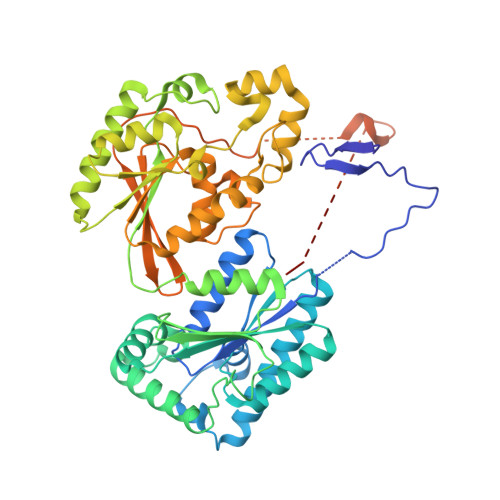
Crystal structure of the hypoxia-inducible form of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB3): a possible new target for cancer therapy.
Kim, S.G., Manes, N.P., El-Maghrabi, M.R., Lee, Y.H.(2006) J Biological Chem 281: 2939-2944
- PubMed: 16316985 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M511019200
- Primary Citation Related Structures:
2AXN - PubMed Abstract:
The hypoxia-inducible form of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB3) plays a crucial role in the progression of cancerous cells by enabling their glycolytic pathways even under severe hypoxic conditions. To understand its structural architecture and to provide a molecular scaffold for the design of new cancer therapeutics, the crystal structure of the human form was determined. The structure at 2.1 A resolution shows that the overall folding and functional dimerization are very similar to those of the liver (PFKFB1) and testis (PFKFB4) forms, as expected from sequence homology. However, in this structure, the N-terminal regulatory domain is revealed for the first time among the PFKFB isoforms. With a beta-hairpin structure, the N terminus interacts with the 2-Pase domain to secure binding of fructose-6-phosphate to the active pocket, slowing down the release of fructose-6-phosphate from the phosphoenzyme intermediate product complex. The C-terminal regulatory domain is mostly disordered, leaving the active pocket of the fructose-2,6-bisphosphatase domain wide open. The active pocket of the 6-phosphofructo-2-kinase domain has a more rigid conformation, allowing independent bindings of substrates, fructose-6-phosphate and ATP, with higher affinities than other isoforms. Intriguingly, the structure shows an EDTA molecule bound to the fructose-6-phosphate site of the 6-phosphofructo-2-kinase active pocket despite its unfavorable liganding concentration, suggesting a high affinity. EDTA is not removable from the site with fructose-6-P alone but is with both ATP and fructose-6-P or with fructose-2,6-bisphosphate. This finding suggests that a molecule in which EDTA is covalently linked to ADP is a good starting molecule for the development of new cancer-therapeutic molecules.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, Louisiana 70803, USA.
Organizational Affiliation: