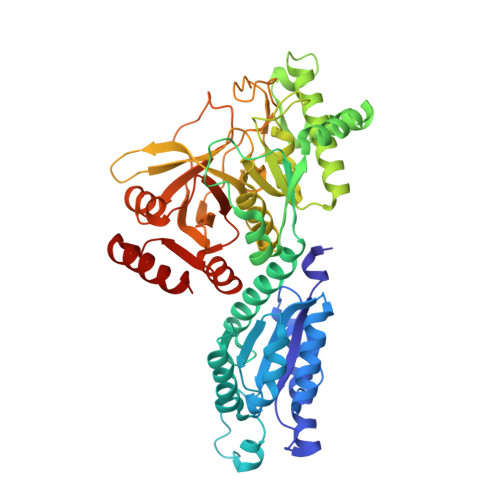
Crystal Structure of Escherichia coli L-Arabinose Isomerase (ECAI), The Putative Target of Biological Tagatose Production
Manjasetty, B.A., Chance, M.R.(2006) J Mol Biology 360: 297-309
- PubMed: 16756997 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.04.040
- Primary Citation Related Structures:
2AJT - PubMed Abstract:
Escherichia coli L-arabinose isomerase (ECAI; EC 5.3.1.4) catalyzes the isomerization of L-arabinose to L-ribulose in vivo. This enzyme is also of commercial interest as it catalyzes the conversion of D-galactose to D-tagatose in vitro. The crystal structure of ECAI was solved and refined at 2.6 A resolution. The subunit structure of ECAI is organised into three domains: an N-terminal, a central and a C-terminal domain. It forms a crystallographic trimeric architecture in the asymmetric unit. Packing within the crystal suggests the idea that ECAI can form a hexameric assembly. Previous electron microscopic and biochemical studies supports that ECAI is hexameric in solution. A comparison with other known structures reveals that ECAI adopts a protein fold most similar to E. coli fucose isomerase (ECFI) despite very low sequence identity 9.7%. The structural similarity between ECAI and ECFI with regard to number of domains, overall fold, biological assembly, and active site architecture strongly suggests that the enzymes have functional similarities. Further, the crystal structure of ECAI forms a basis for identifying molecular determinants responsible for isomerization of arabinose to ribulose in vivo and galactose to tagatose in vitro.
- New York Structural Genomix Research Consortium, Center for Synchrotron Biosciences, National Synchrotron Light Source, Brookhaven National Laboratory, Upton, NY 11973, USA.
Organizational Affiliation: