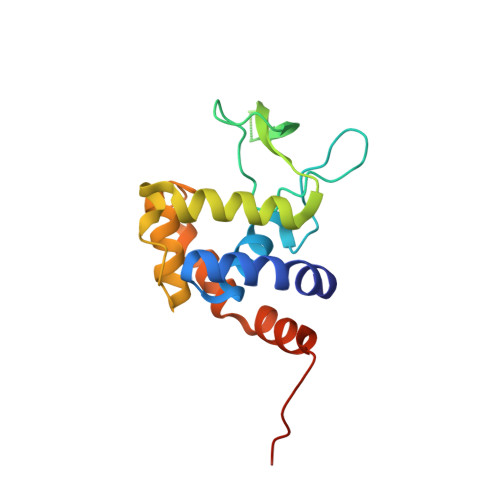
Crystal structure of the glycosidase family 73 peptidoglycan hydrolase FlgJ
Hashimoto, W., Ochiai, A., Momma, K., Itoh, T., Mikami, B., Maruyama, Y., Murata, K.(2009) Biochem Biophys Res Commun 381: 16-21
- PubMed: 19351587 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.01.186
- Primary Citation Related Structures:
2ZYC - PubMed Abstract:
Glycoside hydrolase (GH) categorized into family 73 plays an important role in degrading bacterial cell wall peptidoglycan. The flagellar protein FlgJ contains N- and C-terminal domains responsible for flagellar rod assembly and peptidoglycan hydrolysis, respectively. A member of family GH-73, the C-terminal domain (SPH1045-C) of FlgJ from Sphingomonas sp. strain A1 was expressed in Escherichia coli, purified, and characterized. SPH1045-C exhibited bacterial cell lytic activity most efficiently at pH 6.0 and 37 degrees C. The X-ray crystallographic structure of SPH1045-C was determined at 1.74 A resolution by single-wavelength anomalous diffraction. The enzyme consists of two lobes, alpha and beta. A deep cleft located between the two lobes can accommodate polymer molecules, suggesting that the active site is located in the cleft. Although SPH1045-C shows a structural homology with family GH-22 and GH-23 lysozymes, the arrangement of the nucleophile/base residue in the active site is specific to each peptidoglycan hydrolase.
- Laboratory of Basic and Applied Molecular Biotechnology, Graduate School of Agriculture, Kyoto University, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: