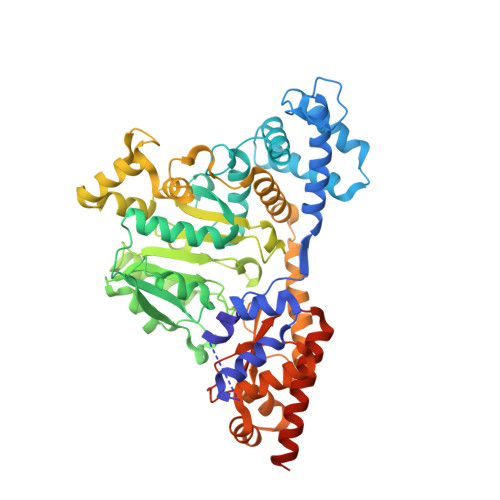
Structure, Assembly, and Mechanism of a PLP-Dependent Dodecameric l-Aspartate beta-Decarboxylase
Chen, H.-J., Ko, T.-P., Lee, C.-Y., Wang, N.-C., Wang, A.H.-J.(2009) Structure 17: 517-529
- PubMed: 19368885 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.02.013
- Primary Citation Related Structures:
2ZY2, 2ZY3, 2ZY4, 2ZY5 - PubMed Abstract:
The type-I PLP enzyme l-aspartate beta-decarboxylase converts aspartate to alanine and CO(2). Similar to the homodimeric aminotransferases, its protein subunit comprises a large and a small domain, of 410 and 120 residues, respectively. The crystal structure reveals a dodecamer made of six identical dimers arranged in a truncated tetrahedron whose assembly involves tetramer and hexamer as intermediates. The additional helical motifs I and II participate in the oligomer formation. Triple mutations of S67R/Y68R/M69R or S67E/Y68E/M69E in motif I produced an inactive dimer. The PLP is bound covalently to Lys315 in the active site, while its phosphate group interacts with a neighboring Tyr134. Removal of the bulky side chain of Arg37, which overhangs the PLP group, improved the substrate affinity. Mutations in flexible regions produced the more active K17A and the completely inactive R487A. The structure also suggests that substrate binding triggers conformational changes essential for catalyzing the reaction.
- Institute of Biological Chemistry, Academia Sinica, Taipei 11529, Taiwan.
Organizational Affiliation: