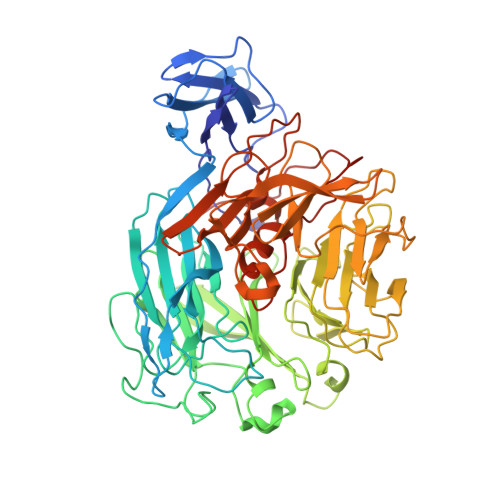
Structural determinants responsible for substrate recognition and mode of action in family 11 polysaccharide lyases
Ochiai, A., Itoh, T., Mikami, B., Hashimoto, W., Murata, K.(2009) J Biological Chem 284: 10181-10189
- PubMed: 19193638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M807799200
- Primary Citation Related Structures:
2ZUX, 2ZUY - PubMed Abstract:
A saprophytic Bacillus subtilis secretes two types of rhamnogalacturonan (RG) lyases, endotype YesW and exotype YesX, which are responsible for an initial cleavage of the RG type I (RG-I) region of plant cell wall pectin. Polysaccharide lyase family 11 YesW and YesX with a significant sequence identity (67.8%) cleave glycoside bonds between rhamnose and galacturonic acid residues in RG-I through a beta-elimination reaction. Here we show the structural determinants for substrate recognition and the mode of action in polysaccharide lyase family 11 lyases. The crystal structures of YesW in complex with rhamnose and ligand-free YesX were determined at 1.32 and 1.65 A resolution, respectively. The YesW amino acid residues such as Asn(152), Asp(172), Asn(532), Gly(533), Thr(534), and Tyr(595) in the active cleft bind to rhamnose molecules through hydrogen bonds and van der Waals contacts. Other rhamnose molecules are accommodated at the noncatalytic domain far from the active cleft, revealing that the domain possibly functions as a novel carbohydrate-binding module. A structural comparison between YesW and YesX indicates that a specific loop in YesX for recognizing the terminal saccharide molecule sterically inhibits penetration of the polymer over the active cleft. The loop-deficient YesX mutant exhibits YesW-like endotype activity, demonstrating that molecular conversion regarding the mode of action is achieved by the addition/removal of the loop for recognizing the terminal saccharide. This is the first report on a structural insight into RG-I recognition and molecular conversion of exotype to endotype in polysaccharide lyases.
- Laboratory of Basic and Applied Molecular Biotechnology.
Organizational Affiliation: