Electron transfer processes within and between proteins containing the HEME C and blue Cu
Nojiri, M., Koteishi, H., Tsuda, A., Nakagami, T., Kobayashi, K., Inoue, T., Yamaguchi, K., Suzuki, S.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Probable nitrite reductase | 442 | Pseudoalteromonas translucida TAC125 | Mutation(s): 0 EC: 1.7.99.3 (PDB Primary Data), 1.7.2.1 (PDB Primary Data) | 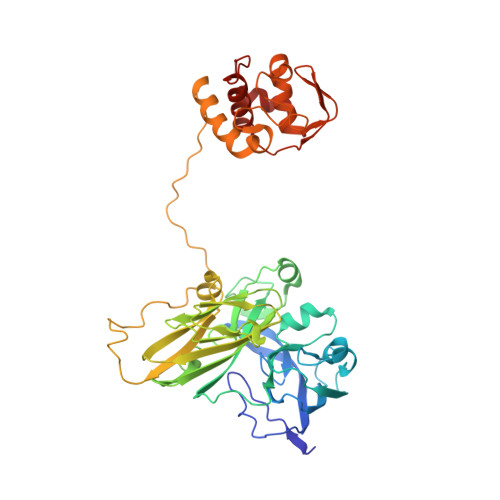 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q3IGF7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | E [auth A] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| SO4 Download:Ideal Coordinates CCD File | F [auth A], G [auth A], H [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| CU Download:Ideal Coordinates CCD File | C [auth A], D [auth A] | COPPER (II) ION Cu JPVYNHNXODAKFH-UHFFFAOYSA-N |  | ||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900003 Query on PRD_900003 | B | sucrose | Oligosaccharide / Nutrient |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 180.338 | α = 90 |
| b = 180.338 | β = 90 |
| c = 180.338 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| SCALEPACK | data scaling |