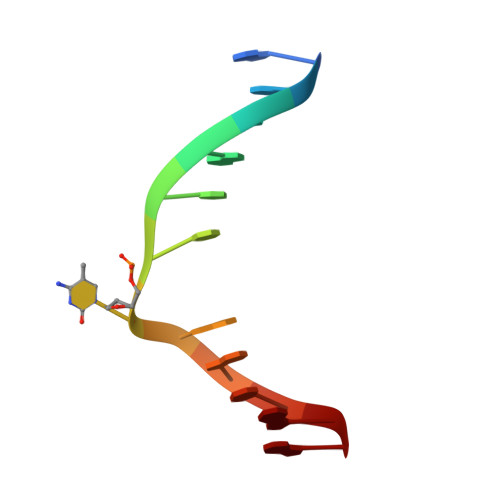
Recognition of hemi-methylated DNA by the SRA protein UHRF1 by a base-flipping mechanism
Arita, K., Ariyoshi, M., Tochio, H., Nakamura, Y., Shirakawa, M.(2008) Nature 455: 818-821
- PubMed: 18772891 Search on PubMed
- DOI: https://doi.org/10.1038/nature07249
- Primary Citation Related Structures:
2ZKD, 2ZKE, 2ZKF, 2ZKG - PubMed Abstract:
DNA methylation of CpG dinucleotides is an important epigenetic modification of mammalian genomes and is essential for the regulation of chromatin structure, of gene expression and of genome stability. Differences in DNA methylation patterns underlie a wide range of biological processes, such as genomic imprinting, inactivation of the X chromosome, embryogenesis, and carcinogenesis. Inheritance of the epigenetic methylation pattern is mediated by the enzyme DNA methyltransferase 1 (Dnmt1), which methylates newly synthesized CpG sequences during DNA replication, depending on the methylation status of the template strands. The protein UHRF1 (also known as Np95 and ICBP90) recognizes hemi-methylation sites via a SET and RING-associated (SRA) domain and directs Dnmt1 to these sites. Here we report the crystal structures of the SRA domain in free and hemi-methylated DNA-bound states. The SRA domain folds into a globular structure with a basic concave surface formed by highly conserved residues. Binding of DNA to the concave surface causes a loop and an amino-terminal tail of the SRA domain to fold into DNA interfaces at the major and minor grooves of the methylation site. In contrast to fully methylated CpG sites recognized by the methyl-CpG-binding domain, the methylcytosine base at the hemi-methylated site is flipped out of the DNA helix in the SRA-DNA complex and fits tightly into a protein pocket on the concave surface. The complex structure suggests that the successive flip out of the pre-existing methylated cytosine and the target cytosine to be methylated is associated with the coordinated transfer of the hemi-methylated CpG site from UHRF1 to Dnmt1.
- Graduate School of Engineering, Kyoto University, Kyoto 615-8510, Japan.
Organizational Affiliation: