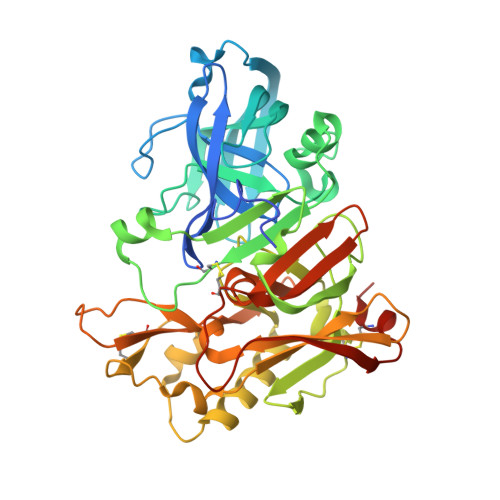
Crystal structure of an active form of BACE1, an enzyme responsible for amyloid beta protein production
Shimizu, H., Tosaki, A., Kaneko, K., Hisano, T., Sakurai, T., Nukina, N.(2008) Mol Cell Biol 28: 3663-3671
- PubMed: 18378702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.02185-07
- Primary Citation Related Structures:
2ZHR, 2ZHS, 2ZHT, 2ZHU, 2ZHV - PubMed Abstract:
BACE1 (beta-secretase) is a transmembrane aspartic protease that cleaves the beta-amyloid precursor protein and generates the amyloid beta peptide (Abeta). BACE1 cycles between the cell surface and the endosomal system many times and becomes activated interconvertibly during its cellular trafficking, leading to the production of Abeta. Here we report the crystal structure of the catalytically active form of BACE1. The active form has novel structural features involving the conformation of the flap and subsites that promote substrate binding. The functionally essential residues and water molecules are well defined and play a key role in the iterative activation of BACE1. We further describe the crystal structure of the dehydrated form of BACE1, showing that BACE1 activity is dependent on the dynamics of a catalytically required Asp-bound water molecule, which directly affects its catalytic properties. These findings provide insight into a novel regulation of BACE1 activity and elucidate how BACE1 modulates its activity during cellular trafficking.
- Laboratory for Structural Neuropathology, RIKEN Brain Science Institute, 2-1 Hirosawa, Wako-shi, Saitama 351-0198, Japan.
Organizational Affiliation: