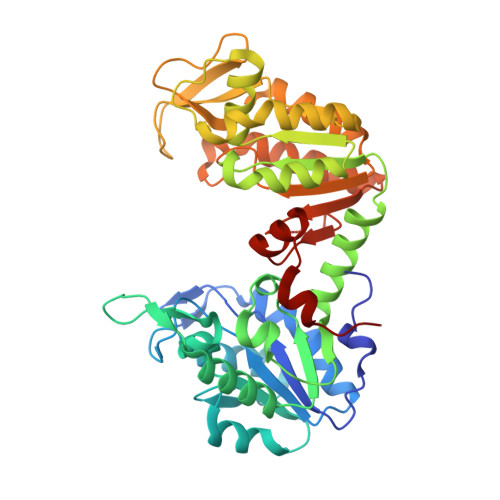
Molecular basis for the lack of enantioselectivity of human 3-phosphoglycerate kinase
Gondeau, C., Chaloin, L., Lallemand, P., Roy, B., Perigaud, C., Barman, T., Varga, A., Vas, M., Lionne, C., Arold, S.T.(2008) Nucleic Acids Res 36: 3620-3629
- PubMed: 18463139 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkn212
- Primary Citation Related Structures:
2ZGV, 3C39, 3C3A, 3C3B, 3C3C - PubMed Abstract:
Non-natural L-nucleoside analogues are increasingly used as therapeutic agents to treat cancer and viral infections. To be active, L-nucleosides need to be phosphorylated to their respective triphosphate metabolites. This stepwise phosphorylation relies on human enzymes capable of processing L-nucleoside enantiomers. We used crystallographic analysis to reveal the molecular basis for the low enantioselectivity and the broad specificity of human 3-phosphoglycerate kinase (hPGK), an enzyme responsible for the last step of phosphorylation of many nucleotide derivatives. Based on structures of hPGK in the absence of nucleotides, and bound to L and d forms of MgADP and MgCDP, we show that a non-specific hydrophobic clamp to the nucleotide base, as well as a water-filled cavity behind it, allows high flexibility in the interaction between PGK and the bases. This, combined with the dispensability of hydrogen bonds to the sugar moiety, and ionic interactions with the phosphate groups, results in the positioning of different nucleotides so to expose their diphosphate group in a position competent for catalysis. Since the third phosphorylation step is often rate limiting, our results are expected to alleviate in silico tailoring of L-type prodrugs to assure their efficient metabolic processing.
- Centre d'études d'agents Pathogènes et Biotechnologies pour la Santé, UMR 5236, CNRS-Universités Montpellier 1 et 2, Institut de Biologie, 4 bd Henri IV, CS69033, 34965 Montpellier cedex 2, France.
Organizational Affiliation: