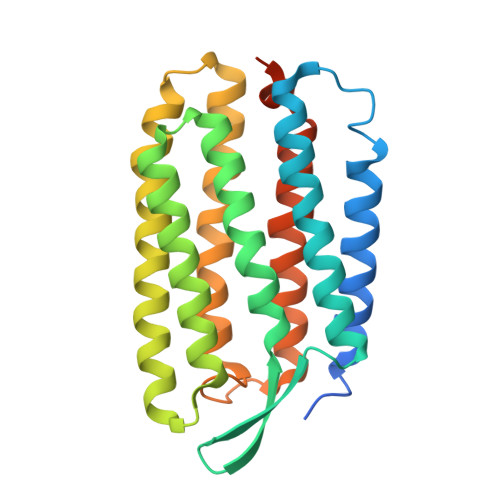
Effect of xenon binding to a hydrophobic cavity on the proton pumping cycle in bacteriorhodopsin
Hayakawa, N., Kasahara, T., Hasegawa, D., Yoshimura, K., Murakami, M., Kouyama, T.(2008) J Mol Biology 384: 812-823
- PubMed: 18930734 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.09.075
- Primary Citation Related Structures:
2ZFE - PubMed Abstract:
To understand the functional role of apolar cavities in bacteriorhodopsin, a light-driven proton pump found in Halobacterium salinarum, we investigated the crystal structure in pressurized xenon or krypton. Diffraction data from the P622 crystal showed that one Xe or Kr atom binds to a preexisting hydrophobic cavity buried between helices C and D, located at the same depth from the membrane surface as Asp96, a key residue in the proton uptake pathway. The occupation fraction of Xe or Kr was calculated as approximately 0.32 at a pressure of 1 MPa. In the unphotolyzed state, the binding of Xe or Kr caused no large deformation of the cavity. However, the proton pumping cycle was greatly perturbed when an aqueous suspension of purple membrane was pressurized with xenon gas; that is, the decay of the M state was accelerated significantly (~5 times at full occupancy), while the decay of an equilibrium state of N and O was slightly decelerated. A similar but much smaller perturbation in the reaction kinetics was observed upon pressurization with krypton gas. In a glycerol/water mixture, xenon-induced acceleration of M decay became less significant in proportion to the water activity. Together with the structure of the xenon-bound protein, these observations suggest that xenon binding helps water molecules permeate into apolar cavities in the proton uptake pathway, thereby accelerating the water-mediated proton transfer from Asp96 to the Schiff base.
- Department of Physics, Graduate School of Science, Nagoya University, Nagoya, Japan.
Organizational Affiliation: