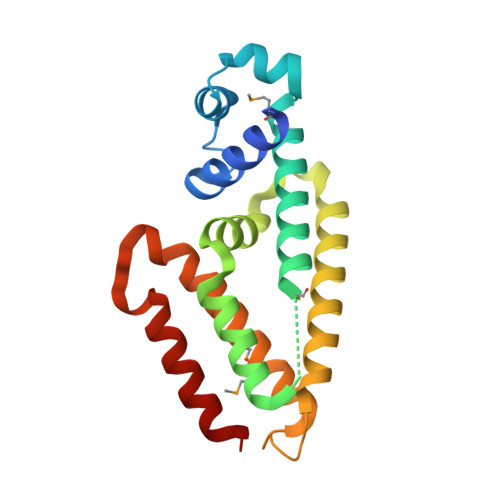
Crystal structure of IcaR, a repressor of the TetR family implicated in biofilm formation in Staphylococcus epidermidis
Jeng, W.Y., Ko, T.P., Liu, C.I., Guo, R.T., Liu, C.L., Shr, H.L., Wang, A.H.J.(2008) Nucleic Acids Res 36: 1567-1577
- PubMed: 18208836 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkm1176
- Primary Citation Related Structures:
2ZCM, 2ZCN - PubMed Abstract:
Expression of the gene cluster icaADBC is necessary for biofilm production in Staphylococcus epidermidis. The ica operon is negatively controlled by the repressor IcaR. Here, the crystal structure of IcaR was determined and the refined structure revealed a homodimer comprising entirely alpha-helices, typical of the tetracycline repressor protein family for gene regulations. The N-terminal domain contains a conserved helix-turn-helix DNA-binding motif with some conformational variations, indicating flexibility in this region. The C-terminal domain shows a complementary surface charge distribution about the dyad axis, ideal for efficient and specific dimer formation. The results of the electrophoretic mobility shift assay and isothermal titration calorimetry suggested that a 28 bp core segment of the ica operator is implicated in the cooperative binding of two IcaR dimers on opposite sides of the duplex DNA. Computer modeling based on the known DNA-complex structure of QacR and site-specific mutagenesis experiments showed that direct protein-DNA interactions are mostly conserved, but with slight variations for recognizing the different sequences. By interfering with the binding of IcaR to DNA, aminoglycoside gentamicin and other antibiotics may activate the icaADBC genes and elicit biofilm production in S. epidermidis, and likely S. aureus, as a defense mechanism.
- Institute of Biological Chemistry, Taipei, Taiwan.
Organizational Affiliation: