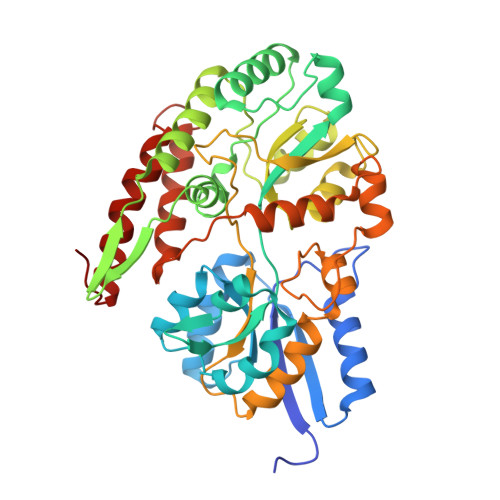
Structural and thermodynamic analyses of solute-binding Protein from Bifidobacterium longum specific for core 1 disaccharide and lacto-N-biose I.
Suzuki, R., Wada, J., Katayama, T., Fushinobu, S., Wakagi, T., Shoun, H., Sugimoto, H., Tanaka, A., Kumagai, H., Ashida, H., Kitaoka, M., Yamamoto, K.(2008) J Biological Chem 283: 13165-13173
- PubMed: 18332142 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M709777200
- Primary Citation Related Structures:
2Z8D, 2Z8E, 2Z8F - PubMed Abstract:
Recently, a gene cluster involving a phosphorylase specific for lacto-N-biose I (LNB; Galbeta1-3GlcNAc) and galacto-N-biose (GNB; Galbeta1-3GalNAc) has been found in Bifidobacterium longum. We showed that the solute-binding protein of a putative ATP-binding cassette-type transporter encoded in the cluster crystallizes only in the presence of LNB or GNB, and therefore we named it GNB/LNB-binding protein (GL-BP). Isothermal titration calorimetry measurements revealed that GL-BP specifically binds LNB and GNB with K(d) values of 0.087 and 0.010 microm, respectively, and the binding process is enthalpy-driven. The crystal structures of GL-BP complexed with LNB, GNB, and lacto-N-tetraose (Galbeta1-3GlcNAcbeta1-3Galbeta1-4Glc) were determined. The interactions between GL-BP and the disaccharide ligands mainly occurred through water-mediated hydrogen bonds. In comparison with the LNB complex, one additional hydrogen bond was found in the GNB complex. These structural characteristics of ligand binding are in agreement with the thermodynamic properties. The overall structure of GL-BP was similar to that of maltose-binding protein; however, the mode of ligand binding and the thermodynamic properties of these proteins were significantly different.
- Department of Biotechnology, the University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: