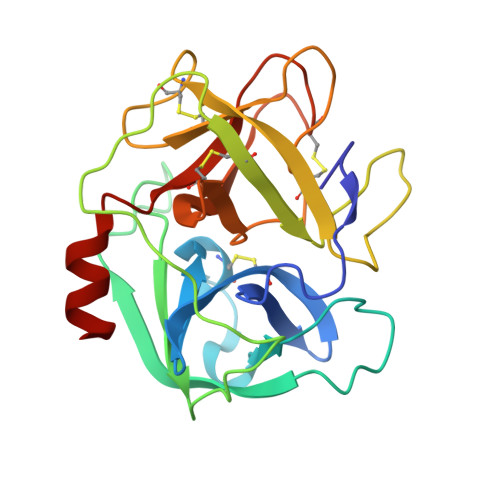
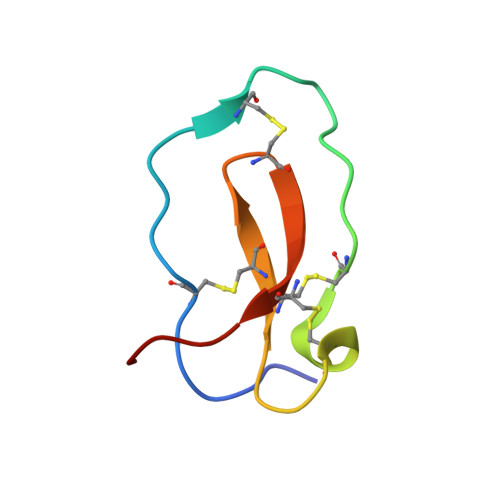
Complex of human neutrophil elastase with 1/2SLPI
Koizumi, M., Fujino, A., Fukushima, K., Kamimura, T., Takimoto-Kamimura, M.(2008) J Synchrotron Radiat 15: 308-311
- PubMed: 18421166
- DOI: https://doi.org/10.1107/S0909049507060670
- Primary Citation of Related Structures:
2Z7F - PubMed Abstract:
SLPI (secretory leukocyte protease inhibitor) is a 107-residue non-glycosylated protease inhibitor, which inhibits a wide range of serine proteases, trypsin, chymotrypsin, neutrophil elastase, chymase and cathepsin G. X-ray crystallographic analyses have shown that SLPI comprises two separate domains of similar architecture [Grütter, Fendrich, Huber & Bode (1988), EMBO J. 7, 345-351] and the C-terminal domain interacts with bovine alpha-chymotrypsin. In order to understand SLPI's multiple functions against various serine proteases, the complex HNE (human neutrophil elastase) has been co-crystallized with 1/2SLPI (recombinant C-terminal domain of SLPI; Arg58-Ala107), which has a biological activity similar to full SLPI. The 1/2SLPI and HNE complex structure was solved at 1.7 A resolution, and compared with the interaction mechanism of elafin, which is a specific inhibitor of elastase. It was found that P1 Leu72i and six hydrogen bonds between the main chains in the primary contact region have sufficient ability to inhibit HNE and PPE (porcine pancreatic elastase), and P5 Tyr68i is important in increasing the selectivity of 1/2SLPI against HNE. The mechanisms of the functions of SLPI are relatively unknown, but the current study could help understand the selectivity of SLPI against HNE and PPE.
- Teijin Institute for Bio-medical Research, Asahigaoka, Hino, Tokyo 191-8512, Japan.
Organizational Affiliation: