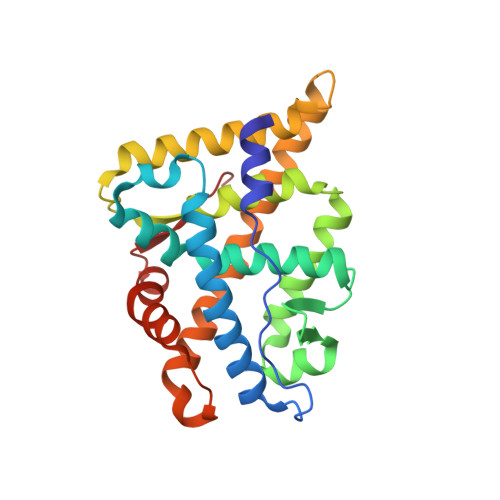
Interaction between the androgen receptor and a segment of its corepressor SHP
Jouravel, N., Sablin, E., Arnold, L.A., Guy, R.K., Fletterick, R.J.(2007) Acta Crystallogr D Biol Crystallogr 63: 1198-1200
- PubMed: 18007036 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444907045702
- Primary Citation Related Structures:
2Z4J - PubMed Abstract:
The mechanisms of functional repression of the androgen receptor (AR) are crucial for the regulation of genes involved in physiological development as well as for the progression of prostate cancer. To date, only two in vivo inhibitors of AR-mediated transcription have been identified: DAX-1 and SHP (small heterodimer partner). SHP is a regulatory nuclear receptor (NR) that lacks DNA-binding and activation domains. Using X-ray crystallography, the interaction between peptide segments of the SHP repressor containing LxxLL-like motifs and the ligand-binding domain of AR have been investigated. Under the crystallization conditions used, it was found that of the three NR Boxes present in the SHP protein sequence, only NR Box 2 (LKKIL motif) formed a complex with AR. Determination of the crystal structure revealed that ten amino acids of the SHP peptide (14-mer) are ordered through interactions with AR. Two side chains make unique interactions that were not reported for other AR-peptide complexes. The NR Box 2 of SHP binds to an adaptable hydrophobic groove on the surface of AR in a fashion observed for other NR-LxxLL-like complexes. Comparisons of AR structures bound to coactivator peptides and the SHP peptide revealed structural similarity of their binding sites, suggesting that transcriptional AR activity may be inhibited by SHP by competing with AR coactivators.
- Department of Biochemistry and Biophysics, University of California, San Francisco, CA 94143, USA.
Organizational Affiliation: