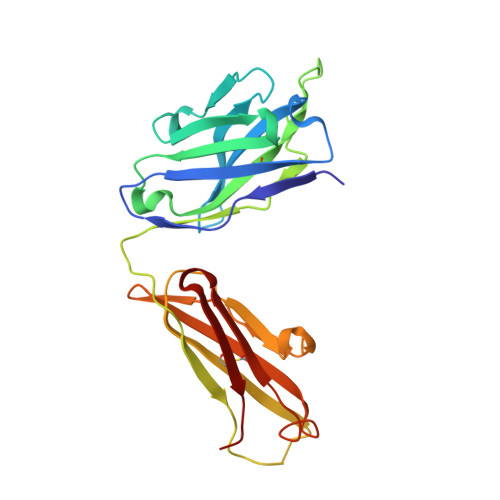
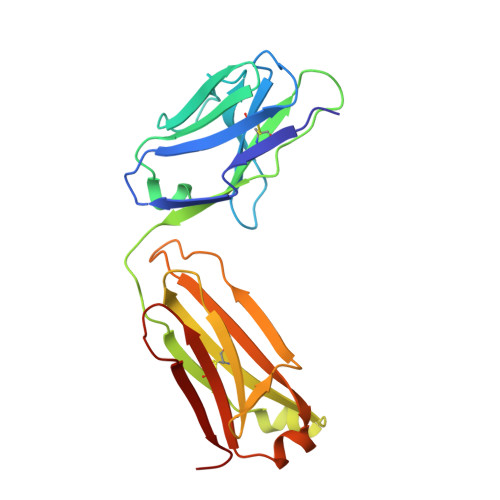
Molecular Characterization of Monoclonal Antibodies that Inhibit Acetylcholinesterase by Targeting the Peripheral Site and Backdoor Region
Bourne, Y., Renault, L., Essono, S., Mondielli, G., Lamourette, P., Boquet, D., Grassi, J., Marchot, P.(2013) PLoS One 8: 77266
- PubMed: 24146971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0077226
- Primary Citation Related Structures:
2YMX - PubMed Abstract:
The inhibition properties and target sites of monoclonal antibodies (mAbs) Elec403, Elec408 and Elec410, generated against Electrophorus electricus acetylcholinesterase (AChE), have been defined previously using biochemical and mutagenesis approaches. Elec403 and Elec410, which bind competitively with each other and with the peptidic toxin inhibitor fasciculin, are directed toward distinctive albeit overlapping epitopes located at the AChE peripheral anionic site, which surrounds the entrance of the active site gorge. Elec408, which is not competitive with the other two mAbs nor fasciculin, targets a second epitope located in the backdoor region, distant from the gorge entrance. To characterize the molecular determinants dictating their binding site specificity, we cloned and sequenced the mAbs; generated antigen-binding fragments (Fab) retaining the parental inhibition properties; and explored their structure-function relationships using complementary x-ray crystallography, homology modeling and flexible docking approaches. Hypermutation of one Elec403 complementarity-determining region suggests occurrence of antigen-driven selection towards recognition of the AChE peripheral site. Comparative analysis of the 1.9Å-resolution structure of Fab408 and of theoretical models of its Fab403 and Fab410 congeners evidences distinctive surface topographies and anisotropic repartitions of charges, consistent with their respective target sites and inhibition properties. Finally, a validated, data-driven docking model of the Fab403-AChE complex suggests a mode of binding at the PAS that fully correlates with the functional data. This comprehensive study documents the molecular peculiarities of Fab403 and Fab410, as the largest peptidic inhibitors directed towards the peripheral site, and those of Fab408, as the first inhibitor directed toward the backdoor region of an AChE and a unique template for the design of new, specific modulators of AChE catalysis.
- Architecture et Fonction des Macromolécules Biologiques (AFMB), CNRS/Aix-Marseille Université, Campus Luminy, Marseille, France.
Organizational Affiliation: