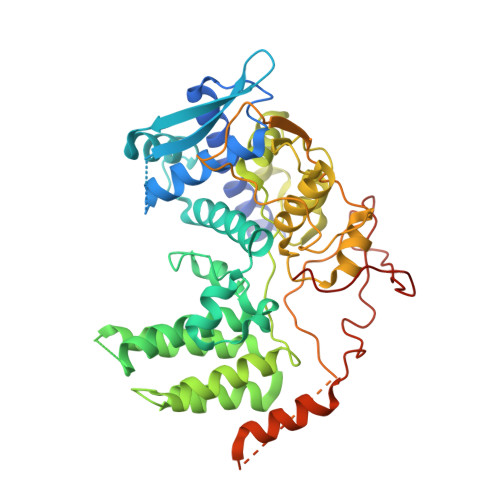
Organization of the Influenza Virus Replication Machinery.
Moeller, A., Kirchdoerfer, R.N., Potter, C.S., Carragher, B., Wilson, I.A.(2012) Science 338: 1631
- PubMed: 23180774
- DOI: https://doi.org/10.1126/science.1227270
- Primary Citation Related Structures:
2YMN - PubMed Abstract:
Influenza virus ribonucleoprotein complexes (RNPs) are central to the viral life cycle and in adaptation to new host species. RNPs are composed of the viral genome, viral polymerase, and many copies of the viral nucleoprotein. In vitro cell expression of all RNP protein components with four of the eight influenza virus gene segments enabled structural determination of native influenza virus RNPs by means of cryogenic electron microscopy (cryo-EM). The cryo-EM structure reveals the architecture and organization of the native RNP, defining the attributes of its largely helical structure and how polymerase interacts with nucleoprotein and the viral genome. Observations of branched-RNP structures in negative-stain electron microscopy and their putative identification as replication intermediates suggest a mechanism for viral replication by a second polymerase on the RNP template.
- National Resource for Automated Molecular Microscopy, Department of Cell Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: