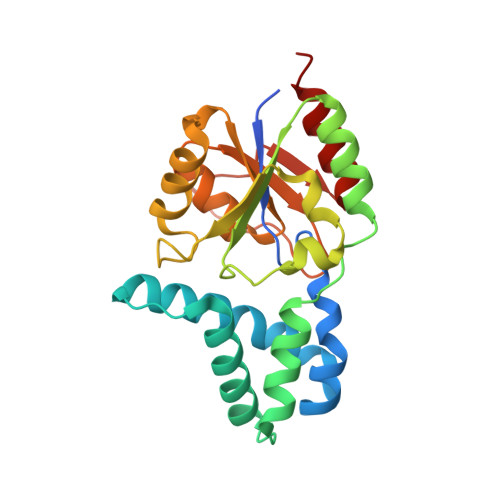
Marine Rhodobacteraceae L-Haloacid Dehalogenase Contains a Novel His/Glu Dyad that Could Activate the Catalytic Water.
Novak, H.R., Sayer, C., Isupov, M.N., Paszkiewicz, K., Gotz, D., Mearns Spragg, A., Littlechild, J.A.(2013) FEBS J 280: 1664
- PubMed: 23384397 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12177
- Primary Citation Related Structures:
2YML, 2YMM, 2YMP, 2YMQ, 2YN4 - PubMed Abstract:
The putative L-haloacid dehalogenase gene (DehRhb) from a marine Rhodobacteraceae family was cloned and overexpressed in Escherichia coli. The DehRhb protein was shown to be an L-haloacid dehalogenase with highest activity towards brominated substrates with short carbon chains (≤ C3). The optimal temperature for enzyme activity was 55 °C, and the Vmax and Km were 1.75 μm·min(-1) ·mg(-1) of protein and 6.72 mm, respectively, when using monobromoacetic acid as a substrate. DehRhb showed moderate thermal stability, with a melting temperature of 67 °C. The enzyme demonstrated high tolerance to solvents, as shown by thermal shift experiments and solvent incubation assays. The DehRhb protein was crystallized and structures of the native, reaction intermediate and substrate-bound forms were determined. The active site of DehRhb had significant differences from previously studied L-haloacid dehalogenases. The asparagine and arginine residues shown to be essential for catalytic activity in other L-haloacid dehalogenases are not present in DehRhb. The histidine residue which replaces the asparagine residue in DehRhb was coordinated by a conformationally strained glutamate residue that replaces a conserved glycine. The His/Glu dyad is positioned for deprotonation of the catalytic water which attacks the ester bond in the reaction intermediate. The catalytic water in DehRhb is shifted by ~ 1.5 Å from its position in other L-haloacid dehalogenases. A similar His/Glu or Asp dyad is known to activate the catalytic water in haloalkane dehalogenases. The DehRhb enzyme represents a novel member within the L-haloacid dehalogenase family and it has potential to be used as a commercial biocatalyst.
- The Henry Wellcome Building for Biocatalysis, Biosciences, College of Life and Environmental Sciences, University of Exeter, Exeter, UK.
Organizational Affiliation: