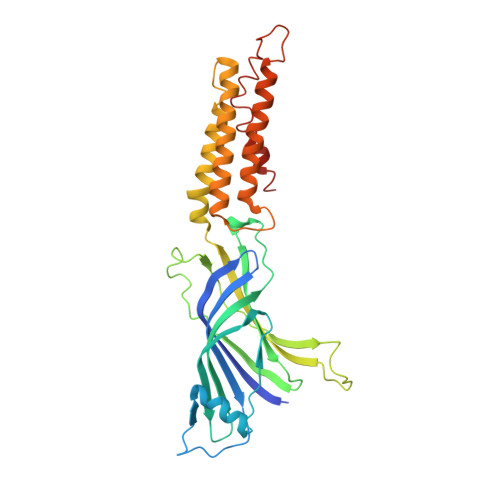
Ligand Activation of the Prokaryotic Pentameric Ligand-Gated Ion Channel Elic.
Zimmermann, I., Dutzler, R.(2011) PLoS Biol 9: 1101
- PubMed: 21713033
- DOI: https://doi.org/10.1371/journal.pbio.1001101
- Primary Citation Related Structures:
2YKS - PubMed Abstract:
While the pentameric ligand-gated ion channel ELIC has recently provided first insight into the architecture of the family at high resolution, its detailed investigation was so far prevented by the fact that activating ligands were unknown. Here we describe a study on the functional characterization of ELIC by electrophysiology and X-ray crystallography. ELIC is activated by a class of primary amines that include the neurotransmitter GABA at high micro- to millimolar concentrations. The ligands bind to a conserved site and evoke currents that slowly desensitize over time. The protein forms cation selective channels with properties that resemble the nicotinic acetylcholine receptor. The high single channel conductance and the comparably simple functional behavior make ELIC an attractive model system to study general mechanisms of ion conduction and gating in this important family of neurotransmitter receptors.
- Department of Biochemistry, University of Zurich, Zurich, Switzerland.
Organizational Affiliation: