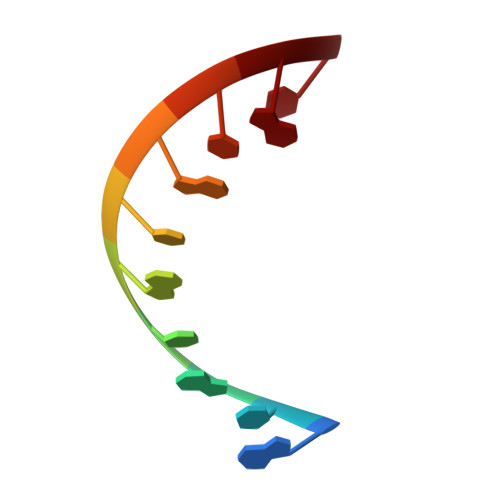
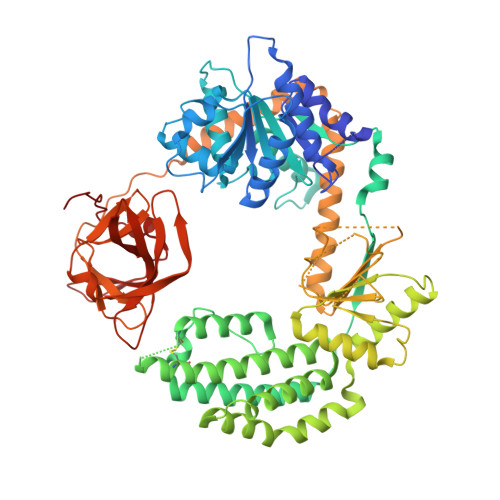
Structural Insights Into RNA Recognition by Rig-I.
Luo, D., Ding, S.C., Vela, A., Kohlway, A., Lindenbach, B.D., Pyle, A.M.(2011) Cell 147: 409
- PubMed: 22000018 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2011.09.023
- Primary Citation Related Structures:
2YKG, 4BPB - PubMed Abstract:
Intracellular RIG-I-like receptors (RLRs, including RIG-I, MDA-5, and LGP2) recognize viral RNAs as pathogen-associated molecular patterns (PAMPs) and initiate an antiviral immune response. To understand the molecular basis of this process, we determined the crystal structure of RIG-I in complex with double-stranded RNA (dsRNA). The dsRNA is sheathed within a network of protein domains that include a conserved "helicase" domain (regions HEL1 and HEL2), a specialized insertion domain (HEL2i), and a C-terminal regulatory domain (CTD). A V-shaped pincer connects HEL2 and the CTD by gripping an α-helical shaft that extends from HEL1. In this way, the pincer coordinates functions of all the domains and couples RNA binding with ATP hydrolysis. RIG-I falls within the Dicer-RIG-I clade of the superfamily 2 helicases, and this structure reveals complex interplay between motor domains, accessory mechanical domains, and RNA that has implications for understanding the nanomechanical function of this protein family and other ATPases more broadly.
- Department of Molecular, Cellular, and Developmental Biology, Yale University, New Haven, CT 06520, USA.
Organizational Affiliation: