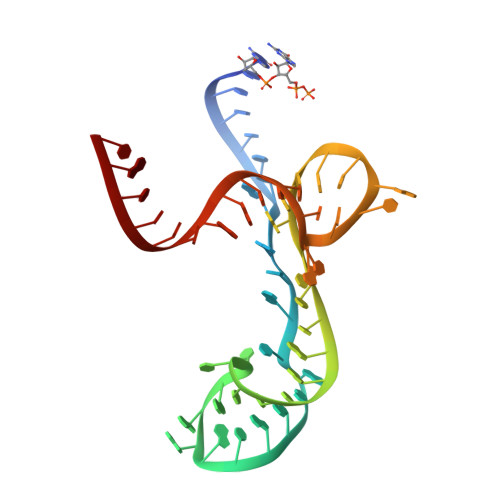
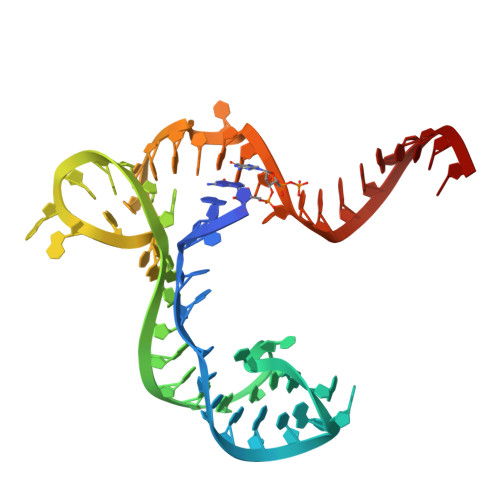
Molecular Sensing by the Aptamer Domain of the Fmn Riboswitch: A General Model for Ligand Binding by Conformational Selection
Vicens, Q., Mondragon, E., Batey, R.T.(2011) Nucleic Acids Res 39: 8586
- PubMed: 21745821 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkr565
- Primary Citation Related Structures:
2YIE, 2YIF - PubMed Abstract:
Understanding the nature of the free state of riboswitch aptamers is important for illuminating common themes in gene regulation by riboswitches. Prior evidence indicated the flavin mononucleotide (FMN)-binding riboswitch aptamer adopted a 'bound-like' structure in absence of FMN, suggesting only local conformational changes upon ligand binding. In the scope of pinpointing the general nature of such changes at the nucleotide level, we performed SHAPE mapping experiments using the aptamer domain of two phylogenetic variants, both in absence and in presence of FMN. We also solved the crystal structures of one of these domains both free (3.3 Å resolution) and bound to FMN (2.95 Å resolution). Our comparative study reveals that structural rearrangements occurring upon binding are restricted to a few of the joining regions that form the binding pocket in both RNAs. This type of binding event with minimal structural perturbations is reminiscent of binding events by conformational selection encountered in other riboswitches and various RNAs.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder, CO 80309-0215, USA.
Organizational Affiliation: