Endothelium-Protective Sphingosine-1-Phosphate Provided by Hdl-Associated Apolipoprotein M.
Christoffersen, C., Obinata, H., Kumaraswamy, S.B., Galvani, S., Ahnstrom, J., Sevvana, M., Egerer-Sieber, C., Muller, Y.A., Hla, T., Nielsen, L.B., Dahlback, B.(2011) Proc Natl Acad Sci U S A 108: 9613
- PubMed: 21606363 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103187108
- Primary Citation Related Structures:
2YG2 - PubMed Abstract:
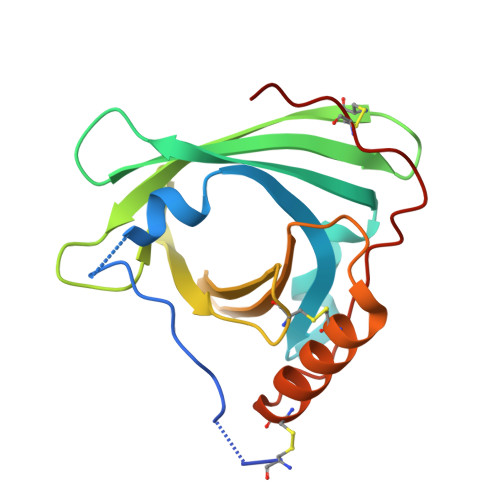
Protection of the endothelium is provided by circulating sphingosine-1-phosphate (S1P), which maintains vascular integrity. We show that HDL-associated S1P is bound specifically to both human and murine apolipoprotein M (apoM). Thus, isolated human ApoM(+) HDL contained S1P, whereas ApoM(-) HDL did not. Moreover, HDL in Apom(-/-) mice contains no S1P, whereas HDL in transgenic mice overexpressing human apoM has an increased S1P content. The 1.7-Å structure of the S1P-human apoM complex reveals that S1P interacts specifically with an amphiphilic pocket in the lipocalin fold of apoM. Human ApoM(+) HDL induced S1P(1) receptor internalization, downstream MAPK and Akt activation, endothelial cell migration, and formation of endothelial adherens junctions, whereas apoM(-) HDL did not. Importantly, lack of S1P in the HDL fraction of Apom(-/-) mice decreased basal endothelial barrier function in lung tissue. Our results demonstrate that apoM, by delivering S1P to the S1P(1) receptor on endothelial cells, is a vasculoprotective constituent of HDL.
- Department of Clinical Biochemistry, Rigshospitalet, 2100 Copenhagen, Denmark.
Organizational Affiliation: