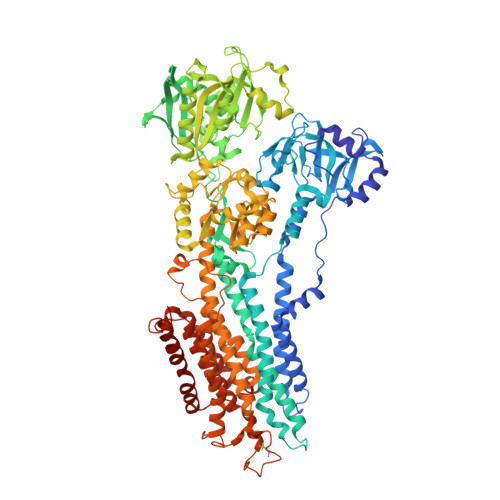
Mutual Adaptation of a Membrane Protein and its Lipid Bilayer During Conformational Changes.
Sonntag, Y., Musgaard, M., Olesen, C., Schiott, B., Moller, J.V., Nissen, P., Thogersen, L.(2011) Nat Commun 2: 304
- PubMed: 21556058 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms1307
- Primary Citation Related Structures:
2YFY - PubMed Abstract:
The structural elucidation of membrane proteins continues to gather pace, but we know little about their molecular interactions with the lipid environment or how they interact with the surrounding bilayer. Here, with the aid of low-resolution X-ray crystallography, we present direct structural information on membrane interfaces as delineated by lipid phosphate groups surrounding the sarco(endo)plasmic reticulum Ca(2+)-ATPase (SERCA) in its phosphorylated and dephosphorylated Ca(2+)-free forms. The protein-lipid interactions are further analysed using molecular dynamics simulations. We find that SERCA adapts to membranes of different hydrophobic thicknesses by inducing local deformations in the lipid bilayers and by undergoing small rearrangements of the amino-acid side chains and helix tilts. These mutually adaptive interactions allow smooth transitions through large conformational changes associated with the transport cycle of SERCA, a strategy that may be of general nature for many membrane proteins.
- Centre for Membrane Pumps in Cells and Disease - PUMPKIN, Danish National Research Foundation, DK-8000 Aarhus C, Denmark.
Organizational Affiliation: