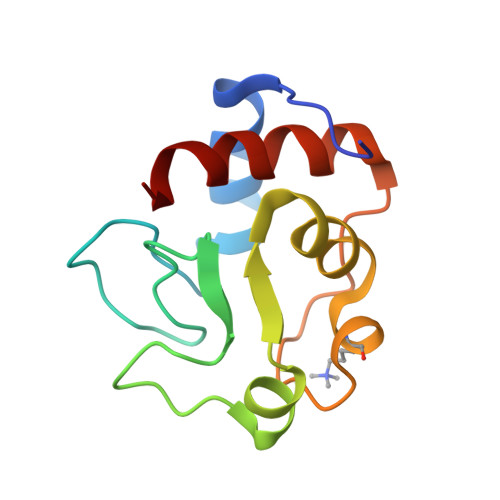
Oxidation state-dependent conformational changes in cytochrome c.
Berghuis, A.M., Brayer, G.D.(1992) J Mol Biology 223: 959-976
- PubMed: 1311391 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90255-i
- Primary Citation Related Structures:
2YCC - PubMed Abstract:
High-resolution three-dimensional structural analyses of yeast iso-1-cytochrome c have now been completed in both oxidation states using isomorphous crystalline material and similar structure determination methodologies. This approach has allowed a comprehensive comparison to be made between these structures and the elucidation of the subtle conformational changes occurring between oxidation states. The structure solution of reduced yeast iso-1-cytochrome c has been published and the determination of the oxidized protein and a comparison of these structures are reported herein. Our data show that oxidation state-dependent changes are expressed for the most part in terms of adjustments to heme structure, movement of internally bound water molecules and segmental thermal parameter changes along the polypeptide chain, rather than as explicit polypeptide chain positional shifts, which are found to be minimal. This result is emphasized by the retention of all main-chain to main-chain hydrogen bond interactions in both oxidation states. Observed thermal factor changes primarily affect four segments of polypeptide chain. Residues 37-39 show less mobility in the oxidized state, with Arg38 and its side-chain being most affected. In contrast, residues 47-59, 65-72 and 81-85 have significantly higher thermal factors, with maximal increases being observed for Asn52, Tyr67 and Phe82. The side-chains of two of these residues are hydrogen bonded to the internally bound water molecule, Wat166, which shows a large 1.7 A displacement towards the positively charged heme iron atom in the oxidized protein. Further analyses suggest that Wat166 is a major factor in stabilizing both oxidation states of the heme through differential orientation of dipole moment, shift in distance to the heme iron atom and alterations in the surrounding hydrogen bonding network. It also seems likely that Wat166 movement leads to the disruption of the hydrogen bond from the side-chain of Tyr67 to the Met80 heme ligand, thereby further stabilizing the positively charged heme iron atom in oxidized cytochrome c. In total, there appear to be three regions about which oxidation state-dependent structural changes are focussed. These include the pyrrole ring A propionate group, Wat166 and the Met80 heme ligand. All three of these foci are linked together by a network of intermediary interactions and are localized to the Met80 ligand side of the heme group. Associated with each is a corresponding nearby segment of polypeptide chain having a substantially higher mobility in the oxidized protein.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Biochemistry, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: