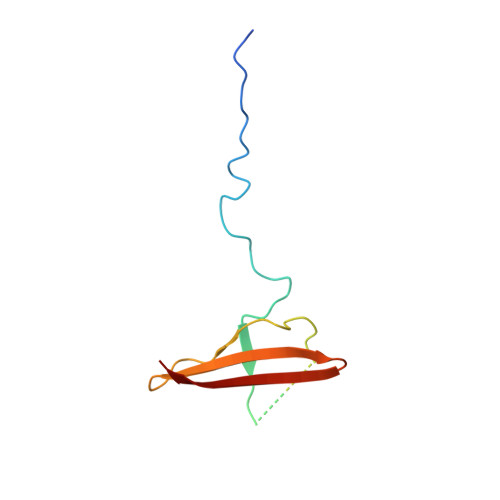
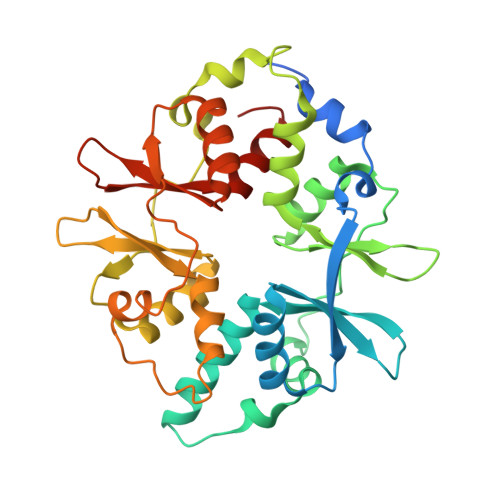
Structure of Mammalian Ampk and its Regulation by Adp
Xiao, B., Sanders, M.J., Underwood, E., Heath, R., Mayer, F., Carmena, D., Jing, C., Walker, P.A., Eccleston, J.F., Haire, L.F., Saiu, P., Howell, S.A., Aasland, R., Martin, S.R., Carling, D., Gamblin, S.J.(2011) Nature 472: 230
- PubMed: 21399626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09932
- Primary Citation Related Structures:
2Y8L, 2Y8Q, 2YA3, 4CFH - PubMed Abstract:
The heterotrimeric AMP-activated protein kinase (AMPK) has a key role in regulating cellular energy metabolism; in response to a fall in intracellular ATP levels it activates energy-producing pathways and inhibits energy-consuming processes. AMPK has been implicated in a number of diseases related to energy metabolism including type 2 diabetes, obesity and, most recently, cancer. AMPK is converted from an inactive form to a catalytically competent form by phosphorylation of the activation loop within the kinase domain: AMP binding to the γ-regulatory domain promotes phosphorylation by the upstream kinase, protects the enzyme against dephosphorylation, as well as causing allosteric activation. Here we show that ADP binding to just one of the two exchangeable AXP (AMP/ADP/ATP) binding sites on the regulatory domain protects the enzyme from dephosphorylation, although it does not lead to allosteric activation. Our studies show that active mammalian AMPK displays significantly tighter binding to ADP than to Mg-ATP, explaining how the enzyme is regulated under physiological conditions where the concentration of Mg-ATP is higher than that of ADP and much higher than that of AMP. We have determined the crystal structure of an active AMPK complex. The structure shows how the activation loop of the kinase domain is stabilized by the regulatory domain and how the kinase linker region interacts with the regulatory nucleotide-binding site that mediates protection against dephosphorylation. From our biochemical and structural data we develop a model for how the energy status of a cell regulates AMPK activity.
- MRC National Institute for Medical Research, The Ridgeway, Mill Hill, London NW7 1AA, UK.
Organizational Affiliation: