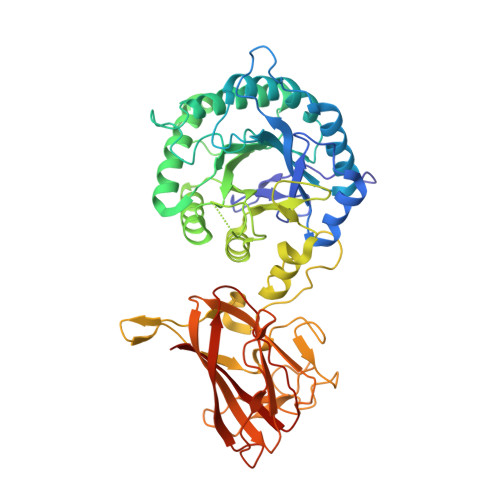
Structure and Function of an Arabinoxylan-Specific Xylanase.
Correia, M.A., Mazumder, K., Bras, J.L., Firbank, S.J., Zhu, Y., Lewis, R.J., York, W.S., Fontes, C.M., Gilbert, H.J.(2011) J Biological Chem 286: 22510
- PubMed: 21378160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.217315
- Primary Citation Related Structures:
2Y8K - PubMed Abstract:
The enzymatic degradation of plant cell walls plays a central role in the carbon cycle and is of increasing environmental and industrial significance. The enzymes that catalyze this process include xylanases that degrade xylan, a β-1,4-xylose polymer that is decorated with various sugars. Although xylanases efficiently hydrolyze unsubstituted xylans, these enzymes are unable to access highly decorated forms of the polysaccharide, such as arabinoxylans that contain arabinofuranose decorations. Here, we show that a Clostridium thermocellum enzyme, designated CtXyl5A, hydrolyzes arabinoxylans but does not attack unsubstituted xylans. Analysis of the reaction products generated by CtXyl5A showed that all the oligosaccharides contain an O3 arabinose linked to the reducing end xylose. The crystal structure of the catalytic module (CtGH5) of CtXyl5A, appended to a family 6 noncatalytic carbohydrate-binding module (CtCBM6), showed that CtGH5 displays a canonical (α/β)(8)-barrel fold with the substrate binding cleft running along the surface of the protein. The catalytic apparatus is housed in the center of the cleft. Adjacent to the -1 subsite is a pocket that could accommodate an l-arabinofuranose-linked α-1,3 to the active site xylose, which is likely to function as a key specificity determinant. CtCBM6, which adopts a β-sandwich fold, recognizes the termini of xylo- and gluco-configured oligosaccharides, consistent with the pocket topology displayed by the ligand-binding site. In contrast to typical modular glycoside hydrolases, there is an extensive hydrophobic interface between CtGH5 and CtCBM6, and thus the two modules cannot function as independent entities.
- Centro de Investigação Interdisciplinar em Sanidade Animal, Faculdade de Medicina Veterinária, Universidade Técnica de Lisboa, Avenida da Universidade Técnica, 1300-477 Lisboa, Portugal.
Organizational Affiliation: